QSMxT#
Author: Ashley Stewart
Date: 25 Aug 2025
License:
Note: If this notebook uses neuroimaging tools from Neurocontainers, those tools retain their original licenses. Please see Neurodesk citation guidelines for details.
Citation and Resources:#
Tools included in this workflow#
QSMxT: Stewart AW, Robinson SD, O’Brien K, et al. QSMxT: Robust masking and artifact reduction for quantitative susceptibility mapping. Magnetic Resonance in Medicine. 2022;87(3):1289-1300. doi:10.1002/mrm.29048
QSMxT: Stewart AW, Bollman S, et al. QSMxT/QSMxT. GitHub; 2022. https://github.com/QSMxT/QSM
Dataset#
Bollmann, S., & Stewart, A. (2023, August 7). QSM DICOM testdata for QSMxT pipeline. Retrieved from osf.io/ru43c
QSMxT Interactive Notebook#
This interactive notebook estimates Quantitative Susceptibility Maps (QSMs) for two gradient-echo (GRE) MRI acquisitions using QSMxT provided by the Neurodesk project.
What is QSM?#
QSM is a form of quantitative MRI (qMRI) that estimates the magnetic susceptibility distribution across an imaged object. Magnetic susceptibility is the degree to which a material becomes magnetised by an external magnetic field. Major contributors to susceptibility include iron, calcium, and myelin, with the susceptibility of water typically approximating a zero-reference, though it is slightly diamagnetic. Read more about QSM here.
What is QSMxT?#
QSMxT is a suite of tools for building and running automated pipelines for QSM that:
is available open-source without any licensing required;
is distributed as a software container making it straightforward to access and install (Neurodesk!)
scales its processing to execute across many acquisitions through jobs parallelisation (using multiple processors or HPCs) provided by Nipype;
automates steps that usually require manual intervention and scripting, including:
DICOM to BIDS conversion;
QSM reconstruction using a range of algorithms;
segmentation using FastSurfer;
group space generation using ANTs;
export of susceptibility statistics by subject and region of interest (ROI) to CSV.
How do I access QSMxT?#
There are a few ways you can access QSMxT:
This notebook: You can access QSMxT in this notebook right now!
If you are running this on a Neurodesk Play instance, you can upload your own data into the sidebar via drag-and-drop.
Neurodesktop: QSMxT is in the applications menu of Neurodesktop.
On Neurodesk Play, upload your own data into the desktop via drag-and-drop.
On a local install of Neurodesk, bring any necessary files into the shared
~/neurodesktop-storagedirectory
Local install: QSMxT can also be installed via the Docker container
HPC install: QSMxT can also be installed via the Singularity container for use on HPCs
Download example DICOMs#
Here, we download some example DICOMs from our OSF repository for QSMxT.
These data include GRE and T1-weighted acquisitions for one subject (duplicated to act as two subjects).
!osf -p ru43c clone . > /dev/null 2>&1
!tar xf osfstorage/dicoms-unsorted.tar
!rm -rf osfstorage/
!tree dicoms-unsorted | head
!echo -e "...\nThere are `ls dicoms-unsorted | wc -l` unsorted DICOMs in ./dicoms-unsorted/"
dicoms-unsorted
├── MR.1.1.dcm
├── MR.1.10.dcm
├── MR.1.100.dcm
├── MR.1.101.dcm
├── MR.1.102.dcm
├── MR.1.103.dcm
├── MR.1.104.dcm
├── MR.1.105.dcm
├── MR.1.106.dcm
...
There are 1216 unsorted DICOMs in ./dicoms-unsorted/
Load QSMxT#
To load QSMxT inside a notebook, we can use the available module system:
import module
await module.load('qsmxt/8.1.1')
!qsmxt --version
[INFO]: QSMxT v8.1.1
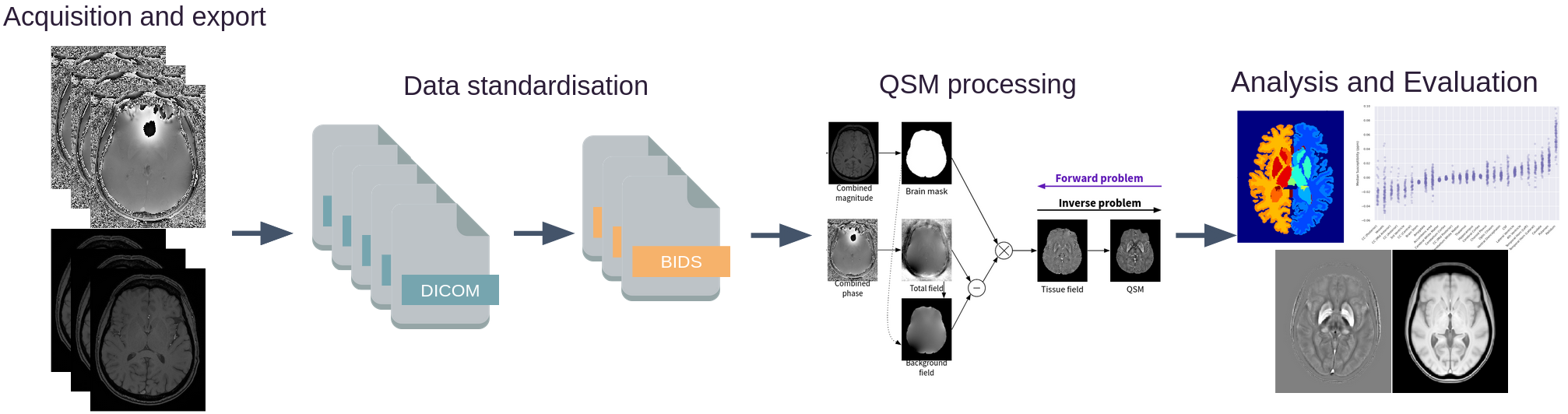
Data standardisation#
QSMxT requires input data to conform to the Brain Imaging Data Structure (BIDS).
Luckily, QSMxT also provides scripts that can convert unorganised NIfTI or DICOM images to BIDS. If you are using NIfTI images and do not have DICOMs, see nifti-convert.
Convert DICOM to BIDS#
The DICOM to BIDS conversion must identify which series should be used for QSM reconstruction (e.g. MEGRE), and which series should be used for segmentation (T1-weighted). Because this information is not stored in the DICOM header in a standardised way, it can only be estimated based on other fields, or the user must identify series themselves.
If QSMxT is run interactively, the user will be prompted to identify the relevant series. However, because we are running QSMxT in a notebook, we disable the interactivity using --auto_yes and allow QSMxT to guess the intended purpose of each acquisition. In this case, it makes the right guess:
!dicom-convert dicoms-unsorted bids --auto_yes
/opt/miniconda-4.12.0/lib/python3.8/site-packages/dicompare/io.py:30: UserWarning: The DICOM readers are highly experimental, unstable, and only work for Siemens time-series at the moment
Please use with caution. We would be grateful for your help in improving them
from nibabel.nicom.csareader import get_csa_header
[INFO]: Running QSMxT 8.1.1
[INFO]: Command: /opt/miniconda-4.12.0/bin/dicom-convert dicoms-unsorted bids --auto_yes
[INFO]: Python interpreter: /opt/miniconda-4.12.0/bin/python3.8
Loading DICOM files: 0%| | 0/1216 [00:00<?, ?it/s]
Loading DICOM files: 0%| | 6/1216 [00:00<00:20, 57.66it/s]
Loading DICOM files: 1%|▏ | 12/1216 [00:00<00:31, 38.50it/s]
Loading DICOM files: 1%|▎ | 17/1216 [00:00<00:28, 42.42it/s]
Loading DICOM files: 2%|▍ | 23/1216 [00:00<00:25, 47.62it/s]
Loading DICOM files: 2%|▍ | 29/1216 [00:00<00:23, 50.56it/s]
Loading DICOM files: 3%|▌ | 37/1216 [00:00<00:20, 57.33it/s]
Loading DICOM files: 4%|▋ | 45/1216 [00:00<00:18, 63.07it/s]
Loading DICOM files: 4%|▉ | 54/1216 [00:00<00:16, 70.01it/s]
Loading DICOM files: 5%|█ | 63/1216 [00:01<00:15, 73.79it/s]
Loading DICOM files: 6%|█▏ | 72/1216 [00:01<00:14, 77.08it/s]
Loading DICOM files: 7%|█▎ | 82/1216 [00:01<00:13, 82.70it/s]
Loading DICOM files: 8%|█▌ | 93/1216 [00:01<00:12, 90.06it/s]
Loading DICOM files: 9%|█▋ | 105/1216 [00:01<00:11, 97.66it/s]
Loading DICOM files: 10%|█▊ | 116/1216 [00:01<00:11, 98.36it/s]
Loading DICOM files: 10%|█▉ | 127/1216 [00:01<00:10, 100.27it/s]
Loading DICOM files: 11%|██ | 138/1216 [00:01<00:10, 101.88it/s]
Loading DICOM files: 12%|██▏ | 149/1216 [00:01<00:10, 102.76it/s]
Loading DICOM files: 13%|██▎ | 160/1216 [00:01<00:10, 100.52it/s]
Loading DICOM files: 14%|██▌ | 171/1216 [00:02<00:10, 100.91it/s]
Loading DICOM files: 15%|██▊ | 182/1216 [00:02<00:10, 99.70it/s]
Loading DICOM files: 16%|██▊ | 194/1216 [00:02<00:09, 102.42it/s]
Loading DICOM files: 17%|███ | 205/1216 [00:02<00:10, 100.48it/s]
Loading DICOM files: 18%|███▍ | 216/1216 [00:02<00:10, 97.65it/s]
Loading DICOM files: 19%|███▌ | 228/1216 [00:02<00:09, 99.97it/s]
Loading DICOM files: 20%|███▌ | 241/1216 [00:02<00:09, 106.84it/s]
Loading DICOM files: 21%|███▋ | 252/1216 [00:02<00:08, 107.20it/s]
Loading DICOM files: 22%|███▉ | 263/1216 [00:02<00:09, 104.43it/s]
Loading DICOM files: 23%|████ | 275/1216 [00:03<00:08, 105.99it/s]
Loading DICOM files: 24%|████▏ | 286/1216 [00:03<00:08, 105.68it/s]
Loading DICOM files: 25%|████▍ | 298/1216 [00:03<00:08, 108.46it/s]
Loading DICOM files: 25%|████▌ | 309/1216 [00:03<00:08, 106.97it/s]
Loading DICOM files: 26%|████▊ | 321/1216 [00:03<00:08, 109.79it/s]
Loading DICOM files: 27%|████▉ | 332/1216 [00:03<00:08, 104.21it/s]
Loading DICOM files: 28%|█████ | 343/1216 [00:03<00:08, 103.73it/s]
Loading DICOM files: 29%|█████▏ | 354/1216 [00:03<00:08, 105.06it/s]
Loading DICOM files: 30%|█████▍ | 365/1216 [00:03<00:08, 102.17it/s]
Loading DICOM files: 31%|█████▌ | 376/1216 [00:04<00:08, 100.99it/s]
Loading DICOM files: 32%|█████▊ | 389/1216 [00:04<00:07, 106.55it/s]
Loading DICOM files: 33%|█████▉ | 400/1216 [00:04<00:07, 105.10it/s]
Loading DICOM files: 34%|██████ | 411/1216 [00:04<00:07, 103.78it/s]
Loading DICOM files: 35%|██████▏ | 422/1216 [00:04<00:07, 103.98it/s]
Loading DICOM files: 36%|██████▍ | 433/1216 [00:04<00:07, 101.77it/s]
Loading DICOM files: 37%|██████▌ | 445/1216 [00:04<00:07, 106.46it/s]
Loading DICOM files: 38%|██████▊ | 457/1216 [00:04<00:06, 108.57it/s]
Loading DICOM files: 39%|██████▉ | 469/1216 [00:04<00:06, 111.18it/s]
Loading DICOM files: 40%|███████ | 481/1216 [00:05<00:06, 113.69it/s]
Loading DICOM files: 41%|███████▎ | 493/1216 [00:05<00:06, 109.91it/s]
Loading DICOM files: 42%|███████▌ | 507/1216 [00:05<00:06, 116.77it/s]
Loading DICOM files: 43%|███████▋ | 520/1216 [00:05<00:05, 119.93it/s]
Loading DICOM files: 44%|███████▉ | 533/1216 [00:05<00:05, 114.36it/s]
Loading DICOM files: 45%|████████ | 545/1216 [00:05<00:05, 113.46it/s]
Loading DICOM files: 46%|████████▏ | 557/1216 [00:05<00:06, 109.23it/s]
Loading DICOM files: 47%|████████▍ | 568/1216 [00:05<00:05, 108.91it/s]
Loading DICOM files: 48%|████████▌ | 579/1216 [00:05<00:06, 103.95it/s]
Loading DICOM files: 49%|████████▋ | 590/1216 [00:06<00:05, 104.60it/s]
Loading DICOM files: 50%|████████▉ | 602/1216 [00:06<00:05, 107.59it/s]
Loading DICOM files: 50%|█████████ | 614/1216 [00:06<00:05, 110.82it/s]
Loading DICOM files: 51%|█████████▎ | 626/1216 [00:06<00:05, 109.86it/s]
Loading DICOM files: 52%|█████████▍ | 638/1216 [00:06<00:05, 110.53it/s]
Loading DICOM files: 53%|█████████▌ | 650/1216 [00:06<00:05, 110.21it/s]
Loading DICOM files: 54%|█████████▊ | 662/1216 [00:06<00:05, 108.04it/s]
Loading DICOM files: 55%|█████████▉ | 674/1216 [00:06<00:04, 110.66it/s]
Loading DICOM files: 56%|██████████▏ | 686/1216 [00:06<00:04, 106.61it/s]
Loading DICOM files: 57%|██████████▎ | 697/1216 [00:07<00:04, 107.31it/s]
Loading DICOM files: 58%|██████████▍ | 709/1216 [00:07<00:04, 108.82it/s]
Loading DICOM files: 59%|██████████▋ | 722/1216 [00:07<00:04, 112.35it/s]
Loading DICOM files: 60%|██████████▊ | 734/1216 [00:07<00:04, 109.24it/s]
Loading DICOM files: 61%|███████████ | 746/1216 [00:07<00:04, 111.40it/s]
Loading DICOM files: 62%|███████████▏ | 758/1216 [00:07<00:04, 108.85it/s]
Loading DICOM files: 63%|███████████▍ | 769/1216 [00:07<00:04, 101.80it/s]
Loading DICOM files: 64%|████████████▏ | 780/1216 [00:07<00:05, 85.86it/s]
Loading DICOM files: 65%|████████████▎ | 790/1216 [00:07<00:04, 89.21it/s]
Loading DICOM files: 66%|████████████▌ | 800/1216 [00:08<00:04, 88.50it/s]
Loading DICOM files: 67%|████████████▋ | 812/1216 [00:08<00:04, 95.19it/s]
Loading DICOM files: 68%|████████████▊ | 823/1216 [00:08<00:03, 98.97it/s]
Loading DICOM files: 69%|████████████▎ | 834/1216 [00:08<00:03, 101.18it/s]
Loading DICOM files: 69%|█████████████▏ | 845/1216 [00:08<00:03, 99.00it/s]
Loading DICOM files: 70%|█████████████▍ | 856/1216 [00:08<00:03, 99.83it/s]
Loading DICOM files: 71%|████████████▊ | 867/1216 [00:08<00:03, 100.26it/s]
Loading DICOM files: 72%|████████████▉ | 878/1216 [00:08<00:03, 101.87it/s]
Loading DICOM files: 73%|█████████████▉ | 889/1216 [00:08<00:03, 98.27it/s]
Loading DICOM files: 74%|██████████████ | 899/1216 [00:09<00:03, 98.72it/s]
Loading DICOM files: 75%|██████████████▏ | 910/1216 [00:09<00:03, 99.45it/s]
Loading DICOM files: 76%|█████████████▋ | 923/1216 [00:09<00:02, 105.40it/s]
Loading DICOM files: 77%|█████████████▊ | 934/1216 [00:09<00:02, 105.84it/s]
Loading DICOM files: 78%|█████████████▉ | 945/1216 [00:09<00:02, 106.60it/s]
Loading DICOM files: 79%|██████████████▏ | 956/1216 [00:09<00:02, 101.22it/s]
Loading DICOM files: 80%|██████████████▎ | 968/1216 [00:09<00:02, 105.54it/s]
Loading DICOM files: 81%|██████████████▍ | 979/1216 [00:09<00:02, 103.61it/s]
Loading DICOM files: 81%|██████████████▋ | 991/1216 [00:09<00:02, 106.26it/s]
Loading DICOM files: 82%|██████████████ | 1002/1216 [00:10<00:02, 103.95it/s]
Loading DICOM files: 83%|██████████████▏ | 1013/1216 [00:10<00:01, 102.52it/s]
Loading DICOM files: 84%|██████████████▎ | 1025/1216 [00:10<00:01, 105.61it/s]
Loading DICOM files: 85%|██████████████▍ | 1036/1216 [00:10<00:01, 105.48it/s]
Loading DICOM files: 86%|██████████████▋ | 1047/1216 [00:10<00:01, 104.22it/s]
Loading DICOM files: 87%|██████████████▊ | 1059/1216 [00:10<00:01, 107.33it/s]
Loading DICOM files: 88%|██████████████▉ | 1070/1216 [00:10<00:01, 104.52it/s]
Loading DICOM files: 89%|███████████████ | 1081/1216 [00:10<00:01, 105.22it/s]
Loading DICOM files: 90%|███████████████▎ | 1093/1216 [00:10<00:01, 106.81it/s]
Loading DICOM files: 91%|███████████████▍ | 1105/1216 [00:10<00:01, 108.01it/s]
Loading DICOM files: 92%|███████████████▌ | 1116/1216 [00:11<00:00, 106.24it/s]
Loading DICOM files: 93%|███████████████▊ | 1127/1216 [00:11<00:00, 104.45it/s]
Loading DICOM files: 94%|████████████████▊ | 1138/1216 [00:11<00:00, 99.15it/s]
Loading DICOM files: 94%|████████████████▉ | 1148/1216 [00:11<00:00, 73.38it/s]
Loading DICOM files: 95%|█████████████████▏| 1157/1216 [00:11<00:00, 67.43it/s]
Loading DICOM files: 96%|█████████████████▏| 1165/1216 [00:11<00:00, 64.08it/s]
Loading DICOM files: 96%|█████████████████▎| 1172/1216 [00:12<00:00, 57.31it/s]
Loading DICOM files: 97%|█████████████████▍| 1179/1216 [00:12<00:00, 57.86it/s]
Loading DICOM files: 98%|█████████████████▌| 1186/1216 [00:12<00:00, 58.72it/s]
Loading DICOM files: 98%|█████████████████▋| 1193/1216 [00:12<00:00, 58.57it/s]
Loading DICOM files: 99%|█████████████████▊| 1202/1216 [00:12<00:00, 64.57it/s]
Loading DICOM files: 99%|█████████████████▉| 1209/1216 [00:12<00:00, 57.89it/s]
Loading DICOM files: 100%|██████████████████| 1216/1216 [00:12<00:00, 95.26it/s]
[INFO]: Loaded DICOM session in 13.13 seconds
/opt/miniconda-4.12.0/lib/python3.8/site-packages/dicompare/acquisition.py:144: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
session_df["BaseAcquisition"] = "acq-" + session_df[acquisition_fields].apply(
/opt/miniconda-4.12.0/lib/python3.8/site-packages/dicompare/acquisition.py:195: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
session_df.loc[param_group.index, "AcquisitionSignature"] = signature
/opt/miniconda-4.12.0/lib/python3.8/site-packages/dicompare/acquisition.py:218: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
session_df["RunNumber"] = 1
/opt/miniconda-4.12.0/lib/python3.8/site-packages/dicompare/acquisition.py:301: PerformanceWarning: DataFrame is highly fragmented. This is usually the result of calling `frame.insert` many times, which has poor performance. Consider joining all columns at once using pd.concat(axis=1) instead. To get a de-fragmented frame, use `newframe = frame.copy()`
session_df["Acquisition"] = session_df["AcquisitionSignature"].fillna("Unknown").astype(str)
[INFO]: Auto-assigning initial labels
[INFO]: Auto-assigned selections:
Acquisition ... Description
0 acq-mp2ragehighres0p5isoslab ...
1 acq-qsmp21mmisote20 ...
2 acq-qsmp21mmisote20 ...
[3 rows x 7 columns]
[INFO]: Merging selections into dataframe...
[INFO]: Processing base group: ('dev_siemens^SB', 1, 20170705, 'acq-mp2ragehighres0p5isoslab', 'mp2rage_highRes_0p5iso_slab_UNI-DEN', 1, '1.2.276.0.7230010.3.1.3.0.4480.1677042698.772469', 'NA')
[INFO]: Found 1 unique Types in this group
[INFO]: Processing Type group: T1w with 288 rows
[INFO]: Found multiple DICOM paths for Type 'T1w', grouping by EchoNumber
[INFO]: Processing Echo group: 1.0 with 288 rows
[INFO]: Converting group with 288 rows
[INFO]: Running command: '"dcm2niix" -o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert" -f "temp_output" -z n -m o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert"'
Chris Rorden's dcm2niiX version v1.0.20240202 (JP2:OpenJPEG) (JP-LS:CharLS) GCC11.4.0 x86-64 (64-bit Linux)
Found 288 DICOM file(s)
Convert 288 DICOM as /home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert/temp_output (378x420x288x1)
Conversion required 0.796477 seconds (0.769504 for core code).
[INFO]: Processing base group: ('dev_siemens^SB', 1, 20170705, 'acq-qsmp21mmisote20', 'QSM_p2_1mmIso_TE20', 1, '1.3.12.2.1107.5.2.43.167001.2017070513471948579426649.0.0.0', 'HC1-7')
[INFO]: Found 1 unique Types in this group
[INFO]: Processing Type group: Mag with 160 rows
[INFO]: Found multiple DICOM paths for Type 'Mag', grouping by EchoNumber
[INFO]: Processing Echo group: 1.0 with 160 rows
[INFO]: Converting group with 160 rows
[INFO]: Running command: '"dcm2niix" -o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert" -f "temp_output" -z n -m o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert"'
Chris Rorden's dcm2niiX version v1.0.20240202 (JP2:OpenJPEG) (JP-LS:CharLS) GCC11.4.0 x86-64 (64-bit Linux)
Found 160 DICOM file(s)
Convert 160 DICOM as /home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert/temp_output (224x224x160x1)
Conversion required 0.190887 seconds (0.189837 for core code).
[INFO]: Processing base group: ('dev_siemens^SB', 1, 20170705, 'acq-qsmp21mmisote20', 'QSM_p2_1mmIso_TE20', 1, '1.3.12.2.1107.5.2.43.167001.2017070513471948580526650.0.0.0', 'HC1-7')
[INFO]: Found 1 unique Types in this group
[INFO]: Processing Type group: Phase with 160 rows
[INFO]: Found multiple DICOM paths for Type 'Phase', grouping by EchoNumber
[INFO]: Processing Echo group: 1.0 with 160 rows
[INFO]: Converting group with 160 rows
[INFO]: Running command: '"dcm2niix" -o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert" -f "temp_output" -z n -m o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert"'
Chris Rorden's dcm2niiX version v1.0.20240202 (JP2:OpenJPEG) (JP-LS:CharLS) GCC11.4.0 x86-64 (64-bit Linux)
Found 160 DICOM file(s)
Convert 160 DICOM as /home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert/temp_output_ph (224x224x160x1)
Conversion required 0.145714 seconds (0.145541 for core code).
[INFO]: Processing base group: ('dev_siemens^SB', 2, 20170705, 'acq-mp2ragehighres0p5isoslab', 'mp2rage_highRes_0p5iso_slab_UNI-DEN', 1, 2, 'NA')
[INFO]: Found 1 unique Types in this group
[INFO]: Processing Type group: T1w with 288 rows
[INFO]: Found multiple DICOM paths for Type 'T1w', grouping by EchoNumber
[INFO]: Processing Echo group: 1.0 with 288 rows
[INFO]: Converting group with 288 rows
[INFO]: Running command: '"dcm2niix" -o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert" -f "temp_output" -z n -m o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert"'
Chris Rorden's dcm2niiX version v1.0.20240202 (JP2:OpenJPEG) (JP-LS:CharLS) GCC11.4.0 x86-64 (64-bit Linux)
Found 288 DICOM file(s)
Convert 288 DICOM as /home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert/temp_output (378x420x288x1)
Conversion required 0.876568 seconds (0.869768 for core code).
[INFO]: Processing base group: ('dev_siemens^SB', 2, 20170705, 'acq-qsmp21mmisote20', 'QSM_p2_1mmIso_TE20', 1, 1, 'HC1-7')
[INFO]: Found 1 unique Types in this group
[INFO]: Processing Type group: Mag with 160 rows
[INFO]: Found multiple DICOM paths for Type 'Mag', grouping by EchoNumber
[INFO]: Processing Echo group: 1.0 with 160 rows
[INFO]: Converting group with 160 rows
[INFO]: Running command: '"dcm2niix" -o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert" -f "temp_output" -z n -m o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert"'
Chris Rorden's dcm2niiX version v1.0.20240202 (JP2:OpenJPEG) (JP-LS:CharLS) GCC11.4.0 x86-64 (64-bit Linux)
Found 160 DICOM file(s)
Convert 160 DICOM as /home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert/temp_output (224x224x160x1)
Conversion required 0.219462 seconds (0.218982 for core code).
[INFO]: Processing base group: ('dev_siemens^SB', 2, 20170705, 'acq-qsmp21mmisote20', 'QSM_p2_1mmIso_TE20', 1, 3, 'HC1-7')
[INFO]: Found 1 unique Types in this group
[INFO]: Processing Type group: Phase with 160 rows
[INFO]: Found multiple DICOM paths for Type 'Phase', grouping by EchoNumber
[INFO]: Processing Echo group: 1.0 with 160 rows
[INFO]: Converting group with 160 rows
[INFO]: Running command: '"dcm2niix" -o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert" -f "temp_output" -z n -m o "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert"'
Chris Rorden's dcm2niiX version v1.0.20240202 (JP2:OpenJPEG) (JP-LS:CharLS) GCC11.4.0 x86-64 (64-bit Linux)
Found 160 DICOM file(s)
Convert 160 DICOM as /home/jovyan/workspace/books/examples/quantitative_imaging/bids/temp_convert/temp_output_ph (224x224x160x1)
Conversion required 0.194597 seconds (0.194441 for core code).
[INFO]: Scanning for multi-coil data to merge...
[INFO]: Scanning for multi-coil runs to combine...
[INFO]: Checking for complex or polar data requiring fixing...
Loading NIfTIs: 0%| | 0/46 [00:00<?, ?it/s]
Loading NIfTIs: 100%|██████████████████████████| 46/46 [00:00<00:00, 782.77it/s]
[INFO]: Finished
!tree bids
bids
├── derivatives
│ ├── qsmxt-2026-04-09-013659
│ │ ├── command.txt
│ │ ├── pypeline.log
│ │ ├── qsmxt.log
│ │ ├── references.txt
│ │ ├── settings.json
│ │ ├── sub-1
│ │ │ └── ses-20170705
│ │ │ ├── anat
│ │ │ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-orig_dseg.nii
│ │ │ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-qsm_dseg.nii
│ │ │ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.json
│ │ │ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.nii
│ │ │ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-orig_dseg.nii
│ │ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_dseg.nii
│ │ │ └── extra_data
│ │ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_desc-t1w-to-qsm_transform.mat
│ │ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-t1w-to-qsm_transform.mat
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_qsm-analysis.csv
│ │ └── sub-2
│ │ └── ses-20170705
│ │ ├── anat
│ │ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-orig_dseg.nii
│ │ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-qsm_dseg.nii
│ │ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.json
│ │ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.nii
│ │ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-orig_dseg.nii
│ │ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_dseg.nii
│ │ └── extra_data
│ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_desc-t1w-to-qsm_transform.mat
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-t1w-to-qsm_transform.mat
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_qsm-analysis.csv
│ └── workflow
│ └── qsmxt-workflow
│ ├── d3.js
│ ├── graph.json
│ ├── graph1.json
│ ├── index.html
│ ├── sub-1
│ │ └── ses-20170705
│ │ ├── qsm_acq-acqmp2ragehighres0p5isoslab_T1w
│ │ │ ├── ants_register-t1-to-qsm
│ │ │ │ ├── _0x3b6e48b848b69bcf3143b0e36fc954c9.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ ├── command.txt
│ │ │ │ ├── result_ants_register-t1-to-qsm.pklz
│ │ │ │ └── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_0GenericAffine.mat
│ │ │ ├── ants_transform-segmentation-to-qsm
│ │ │ │ ├── _0xe6d5ec26a855ebfd4717a00b56193369.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ ├── command.txt
│ │ │ │ ├── result_ants_transform-segmentation-to-qsm.pklz
│ │ │ │ └── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_segmentation_trans.nii
│ │ │ ├── copyfile
│ │ │ │ ├── _0xd8c663486cbc60da130fb8d1025dd48e.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ └── result_copyfile.pklz
│ │ │ ├── fastsurfer_segment-t1
│ │ │ │ ├── _0x17109bf62edc298fda0930bd7b2b45fc.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ ├── command.txt
│ │ │ │ ├── output
│ │ │ │ │ ├── mri
│ │ │ │ │ │ └── orig
│ │ │ │ │ └── scripts
│ │ │ │ ├── result_fastsurfer_segment-t1.pklz
│ │ │ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg.mgz
│ │ │ │ └── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg_nii.nii
│ │ │ ├── func_getfirst-canonical-magnitude
│ │ │ │ ├── _0x839d45b873256671bec4a0e0975faa21.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ └── result_func_getfirst-canonical-magnitude.pklz
│ │ │ ├── func_read-json-me
│ │ │ │ ├── _0xc04d90e741d1629635e91861bde200b6.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ └── result_func_read-json-me.pklz
│ │ │ ├── func_read-json-se
│ │ │ │ ├── _0x2f3b1fe946b4ba7ff15bd9bf10bb7325.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ └── result_func_read-json-se.pklz
│ │ │ ├── nibabel_as-canonical
│ │ │ │ ├── _0x90cc5fb5dd6c3ddc32cd3ad1a2c35b68.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ └── result_nibabel_as-canonical.pklz
│ │ │ ├── nibabel_numpy_nilearn_t1w-seg-resampled
│ │ │ │ ├── _0xb40413c9ea64d494f5b22f74588037e1.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ ├── result_nibabel_numpy_nilearn_t1w-seg-resampled.pklz
│ │ │ │ └── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg_nii_resampled.nii
│ │ │ ├── nibabel_read-nii
│ │ │ │ ├── _0x40d0c1e5ec310206ef8c3220e723cd36.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ └── result_nibabel_read-nii.pklz
│ │ │ └── numpy_numpy_nibabel_mgz2nii
│ │ │ ├── _0x2a86300d9ab35d17fd0565eb0033c51d.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_numpy_numpy_nibabel_mgz2nii.pklz
│ │ └── qsm_acq-acqqsmp21mmisote20_T2starw
│ │ ├── ants_register-t1-to-qsm
│ │ │ ├── _0xa1db2b9552f37480d00edb7ab08bf5fc.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── result_ants_register-t1-to-qsm.pklz
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_0GenericAffine.mat
│ │ ├── ants_transform-segmentation-to-qsm
│ │ │ ├── _0x941af1e40bf849698a1261b027139f20.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── result_ants_transform-segmentation-to-qsm.pklz
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_segmentation_trans.nii
│ │ ├── combine_lists1
│ │ │ ├── _0x1de7fa43f8ddd7ab6085b437fc03804c.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_combine_lists1.pklz
│ │ ├── combine_lists2
│ │ │ ├── _0x731e1d78245239382cb32d0a2b76e46e.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_combine_lists2.pklz
│ │ ├── copy_qsm_json_sidecar
│ │ │ ├── _0x24d29ee452153e8280215c9dc4d3bf36.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_copy_qsm_json_sidecar.pklz
│ │ ├── copyfile
│ │ │ ├── _0xf9b09c0f16d17041ac1dce7fa1b0232c.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_copyfile.pklz
│ │ ├── create_permutations
│ │ │ ├── _0x23ddfa1efd8437c385c24be26b9c36f2.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_create_permutations.pklz
│ │ ├── fastsurfer_segment-t1
│ │ │ ├── _0x17109bf62edc298fda0930bd7b2b45fc.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── output
│ │ │ │ ├── mri
│ │ │ │ │ └── orig
│ │ │ │ └── scripts
│ │ │ ├── result_fastsurfer_segment-t1.pklz
│ │ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg.mgz
│ │ │ └── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg_nii.nii
│ │ ├── func_getfirst-canonical-magnitude
│ │ │ ├── _0xc836bd2c385f56260f08db13c9a6e72b.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_func_getfirst-canonical-magnitude.pklz
│ │ ├── func_read-json-me
│ │ │ ├── _0xe09bae5e56a0a28e09f5ad011d160b39.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_func_read-json-me.pklz
│ │ ├── func_read-json-se
│ │ │ ├── _0x2b23e33a017a8bead72aa5d2c0ea3fa7.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_func_read-json-se.pklz
│ │ ├── mask_workflow
│ │ │ ├── fsl-bet
│ │ │ │ ├── _0x2495113342ce59b92d010584d6838ef0.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ ├── command.txt
│ │ │ │ ├── result_fsl-bet.pklz
│ │ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw_bet-mask.nii.gz
│ │ │ └── scipy_numpy_nibabel_bet_erode
│ │ │ ├── _0x171c854689a8dfc03b756ef9c56bbde0.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── result_scipy_numpy_nibabel_bet_erode.pklz
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw_bet-mask_ero.nii.gz
│ │ ├── nibabel_as-canonical
│ │ │ ├── _0xf2b5e5f7ab4a0fad530024a79f7d1072.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_nibabel_as-canonical.pklz
│ │ ├── nibabel_numpy_analyse-qsm
│ │ │ ├── _0x30809269d048221995157233f98854ba.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── mapflow
│ │ │ │ └── _nibabel_numpy_analyse-qsm0
│ │ │ │ ├── _0x361f0330afba5dcacd23fe0ba9a524ff.json
│ │ │ │ ├── _inputs.pklz
│ │ │ │ ├── _node.pklz
│ │ │ │ ├── _report
│ │ │ │ │ └── report.rst
│ │ │ │ ├── result__nibabel_numpy_analyse-qsm0.pklz
│ │ │ │ └── sub-1_ses-20170705_desc-qsmxt-2026-04-09-013659_Chimap_sub-1_ses-20170705_space-qsm_desc-qsmxt-2026-04-09-013659_dseg_analysis.csv
│ │ │ └── result_nibabel_numpy_analyse-qsm.pklz
│ │ ├── nibabel_numpy_nilearn_axial-resampling
│ │ │ ├── _0x914a7bc37844a6588b9268c16df896f4.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_nibabel_numpy_nilearn_axial-resampling.pklz
│ │ ├── nibabel_numpy_nilearn_qsm-resampled
│ │ │ ├── _0x9b48140c1aa9273df6ffffe3a812066a.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_nibabel_numpy_nilearn_qsm-resampled.pklz
│ │ ├── nibabel_numpy_nilearn_t1w-seg-resampled
│ │ │ ├── _0xbea33b67ded5907657f5b3e5931ab351.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── result_nibabel_numpy_nilearn_t1w-seg-resampled.pklz
│ │ │ └── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg_nii_resampled.nii
│ │ ├── nibabel_numpy_qsm-average
│ │ │ ├── _0x1a25f22192ea29532ccd2e899fd1cd6e.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_nibabel_numpy_qsm-average.pklz
│ │ ├── nibabel_numpy_qsm-referenced
│ │ │ ├── _0xe778e57dd40fce2b0bab992a5a67ee29.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── result_nibabel_numpy_qsm-referenced.pklz
│ │ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized_vsharp_rts_ref.json
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized_vsharp_rts_ref.nii
│ │ ├── nibabel_numpy_scale-phase
│ │ │ ├── _0x3cac41a19d9418ce1311fb85ee11843f.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── result_nibabel_numpy_scale-phase.pklz
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled.nii
│ │ ├── nibabel_read-nii
│ │ │ ├── _0xf75026f5ce2974037cfd29db72bb5be8.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_nibabel_read-nii.pklz
│ │ ├── numpy_numpy_nibabel_mgz2nii
│ │ │ ├── _0xba7cc087d8181c0dc26dea8eb57dddf1.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_numpy_numpy_nibabel_mgz2nii.pklz
│ │ └── qsm_workflow
│ │ ├── mrt_romeo
│ │ │ ├── _0xee872193f1e2e309d224f571679c45f2.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── result_mrt_romeo.pklz
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped.nii
│ │ ├── nibabel-numpy_normalize-phase
│ │ │ ├── _0xc2726a427e641aced69023605641defa.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── result_nibabel-numpy_normalize-phase.pklz
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized.nii
│ │ ├── qsmjl_rts
│ │ │ ├── _0x1765da7adfac8eb58b3184797bcf3d20.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── result_qsmjl_rts.pklz
│ │ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized_vsharp_rts.nii
│ │ └── qsmjl_vsharp
│ │ ├── _0x78858b6db141733ae8b84d3a72dcb4dd.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── command.txt
│ │ ├── result_qsmjl_vsharp.pklz
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw_bet-mask_ero_vsharp-mask.nii
│ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized_vsharp.nii
│ └── sub-2
│ └── ses-20170705
│ ├── qsm_acq-acqmp2ragehighres0p5isoslab_T1w
│ │ ├── ants_register-t1-to-qsm
│ │ │ ├── _0xa2d4c01515da0cd570ad9dbce40043d8.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── result_ants_register-t1-to-qsm.pklz
│ │ │ └── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_0GenericAffine.mat
│ │ ├── ants_transform-segmentation-to-qsm
│ │ │ ├── _0x2d24ac26cf585573ec8290d779411df7.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── result_ants_transform-segmentation-to-qsm.pklz
│ │ │ └── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_segmentation_trans.nii
│ │ ├── copyfile
│ │ │ ├── _0x34672f98f88ff68dc67c919d3af21141.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_copyfile.pklz
│ │ ├── fastsurfer_segment-t1
│ │ │ ├── _0x78cb1ca06599da70c905b0a51f8bde07.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── output
│ │ │ │ ├── mri
│ │ │ │ │ └── orig
│ │ │ │ └── scripts
│ │ │ ├── result_fastsurfer_segment-t1.pklz
│ │ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg.mgz
│ │ │ └── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg_nii.nii
│ │ ├── func_getfirst-canonical-magnitude
│ │ │ ├── _0x5dcf11750bbb8aa2aac5857f53cbacbc.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_func_getfirst-canonical-magnitude.pklz
│ │ ├── func_read-json-me
│ │ │ ├── _0xfe0117efa8cdca54e4b789c2f3b667bd.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_func_read-json-me.pklz
│ │ ├── func_read-json-se
│ │ │ ├── _0x2887a489c7bd7f5eaadbe7d7acc57671.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_func_read-json-se.pklz
│ │ ├── nibabel_as-canonical
│ │ │ ├── _0xa141fbd6be4d5af27de181a0b52f1cc4.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_nibabel_as-canonical.pklz
│ │ ├── nibabel_numpy_nilearn_t1w-seg-resampled
│ │ │ ├── _0x68fa5d80f7d47b0dace1634955c919df.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── result_nibabel_numpy_nilearn_t1w-seg-resampled.pklz
│ │ │ └── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg_nii_resampled.nii
│ │ ├── nibabel_read-nii
│ │ │ ├── _0x92abd1a2a550d179979ee543f5ef041a.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ └── result_nibabel_read-nii.pklz
│ │ └── numpy_numpy_nibabel_mgz2nii
│ │ ├── _0x95ca803a131da0662e39bb561be8defd.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_numpy_numpy_nibabel_mgz2nii.pklz
│ └── qsm_acq-acqqsmp21mmisote20_T2starw
│ ├── ants_register-t1-to-qsm
│ │ ├── _0x03fd0fe41bf64322d2c189060448f97a.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── command.txt
│ │ ├── result_ants_register-t1-to-qsm.pklz
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_0GenericAffine.mat
│ ├── ants_transform-segmentation-to-qsm
│ │ ├── _0x98f2d503bc396fcba3c588893fbfb417.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── command.txt
│ │ ├── result_ants_transform-segmentation-to-qsm.pklz
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_segmentation_trans.nii
│ ├── combine_lists1
│ │ ├── _0x895206669c0dea6c514fc0e258ab46c4.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_combine_lists1.pklz
│ ├── combine_lists2
│ │ ├── _0x8024fe0d1f8e0aea35730e6be250805f.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_combine_lists2.pklz
│ ├── copy_qsm_json_sidecar
│ │ ├── _0xd0bddb8ae321e63f65c184465f7bc44d.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_copy_qsm_json_sidecar.pklz
│ ├── copyfile
│ │ ├── _0xe6db5b93339bb7ac6c8cad4b85e2de84.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_copyfile.pklz
│ ├── create_permutations
│ │ ├── _0xd6556ad02e4af5193f0f669d20d1e083.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_create_permutations.pklz
│ ├── fastsurfer_segment-t1
│ │ ├── _0x78cb1ca06599da70c905b0a51f8bde07.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── command.txt
│ │ ├── output
│ │ │ ├── mri
│ │ │ │ └── orig
│ │ │ └── scripts
│ │ ├── result_fastsurfer_segment-t1.pklz
│ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg.mgz
│ │ └── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg_nii.nii
│ ├── func_getfirst-canonical-magnitude
│ │ ├── _0xb201ec012a2fdd0e426cd9d00cb5a202.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_func_getfirst-canonical-magnitude.pklz
│ ├── func_read-json-me
│ │ ├── _0xff64a069d6bdc4e2ade84381fe750a18.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_func_read-json-me.pklz
│ ├── func_read-json-se
│ │ ├── _0x0461a17932975ecfc60c19abf13899d0.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_func_read-json-se.pklz
│ ├── mask_workflow
│ │ ├── fsl-bet
│ │ │ ├── _0x13a6e3a87cdc9d2cc44c6f1fe2485ab3.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── command.txt
│ │ │ ├── result_fsl-bet.pklz
│ │ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw_bet-mask.nii.gz
│ │ └── scipy_numpy_nibabel_bet_erode
│ │ ├── _0x1422f7f78303721d29f734ccbb7f817c.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── result_scipy_numpy_nibabel_bet_erode.pklz
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw_bet-mask_ero.nii.gz
│ ├── nibabel_as-canonical
│ │ ├── _0x12ed99e1d3503a8031080296ada00732.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_nibabel_as-canonical.pklz
│ ├── nibabel_numpy_analyse-qsm
│ │ ├── _0x03a4ca69dd04c221829860002fb714e8.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── mapflow
│ │ │ └── _nibabel_numpy_analyse-qsm0
│ │ │ ├── _0x6e983a06a2be9d52131286bd80b0c358.json
│ │ │ ├── _inputs.pklz
│ │ │ ├── _node.pklz
│ │ │ ├── _report
│ │ │ │ └── report.rst
│ │ │ ├── result__nibabel_numpy_analyse-qsm0.pklz
│ │ │ └── sub-2_ses-20170705_desc-qsmxt-2026-04-09-013659_Chimap_sub-2_ses-20170705_space-qsm_desc-qsmxt-2026-04-09-013659_dseg_analysis.csv
│ │ └── result_nibabel_numpy_analyse-qsm.pklz
│ ├── nibabel_numpy_nilearn_axial-resampling
│ │ ├── _0x45b2e1a8f33f0615cb727f84f2cb1017.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_nibabel_numpy_nilearn_axial-resampling.pklz
│ ├── nibabel_numpy_nilearn_qsm-resampled
│ │ ├── _0x9db84b44c24b4a042e0b99747be64145.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_nibabel_numpy_nilearn_qsm-resampled.pklz
│ ├── nibabel_numpy_nilearn_t1w-seg-resampled
│ │ ├── _0xc2ddaa6299326db61a3ee67295d17231.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── result_nibabel_numpy_nilearn_t1w-seg-resampled.pklz
│ │ └── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w_dseg_nii_resampled.nii
│ ├── nibabel_numpy_qsm-average
│ │ ├── _0x4a03c50a621bc91dc67914e0161f1bed.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_nibabel_numpy_qsm-average.pklz
│ ├── nibabel_numpy_qsm-referenced
│ │ ├── _0x8f01f35ca881b9c5935bbb9566896c65.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── result_nibabel_numpy_qsm-referenced.pklz
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized_vsharp_rts_ref.json
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized_vsharp_rts_ref.nii
│ ├── nibabel_numpy_scale-phase
│ │ ├── _0x9500f7ecd2ced0acfe1846d4db055416.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── result_nibabel_numpy_scale-phase.pklz
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled.nii
│ ├── nibabel_read-nii
│ │ ├── _0x0cd45d1a288c65e4e8558bf62ed8fe4b.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_nibabel_read-nii.pklz
│ ├── numpy_numpy_nibabel_mgz2nii
│ │ ├── _0x652e9c131f23986b0503163682f95153.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ └── result_numpy_numpy_nibabel_mgz2nii.pklz
│ └── qsm_workflow
│ ├── mrt_romeo
│ │ ├── _0x2c072cb526f46db911c658a7be39116d.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── command.txt
│ │ ├── result_mrt_romeo.pklz
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped.nii
│ ├── nibabel-numpy_normalize-phase
│ │ ├── _0x37152edf32a7cc4cc10023c22561948f.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── result_nibabel-numpy_normalize-phase.pklz
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized.nii
│ ├── qsmjl_rts
│ │ ├── _0x5bfadc6b2d87dff442025fdde0013945.json
│ │ ├── _inputs.pklz
│ │ ├── _node.pklz
│ │ ├── _report
│ │ │ └── report.rst
│ │ ├── command.txt
│ │ ├── result_qsmjl_rts.pklz
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized_vsharp_rts.nii
│ └── qsmjl_vsharp
│ ├── _0x5a55e684d9d7f3c4f9aa6578f8dd315a.json
│ ├── _inputs.pklz
│ ├── _node.pklz
│ ├── _report
│ │ └── report.rst
│ ├── command.txt
│ ├── result_qsmjl_vsharp.pklz
│ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw_bet-mask_ero_vsharp-mask.nii
│ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw_scaled_romeo-unwrapped_normalized_vsharp.nii
├── log_2026-04-09_01-36-19030265.txt
├── log_2026-04-09_02-04-50534677.txt
├── references.txt
├── sub-1
│ └── ses-20170705
│ └── anat
│ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w.json
│ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w.nii
│ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw.json
│ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw.nii
│ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw.json
│ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw.nii
└── sub-2
└── ses-20170705
└── anat
├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w.json
├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_part-mag_T1w.nii
├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw.json
├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-mag_T2starw.nii
├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw.json
└── sub-2_ses-20170705_acq-acqqsmp21mmisote20_part-phase_T2starw.nii
205 directories, 494 files
Inspect input data#
Here we define a function we will use to visualise NIfTI images so we can view some of the input data:
%%capture
!pip install seaborn numpy nibabel pandas nilearn
from glob import glob
from matplotlib import pyplot as plt
import numpy as np
import nibabel as nib
def show_nii(nii_path, title=None, figsize=(4,5), cmap='gray', **imshow_args):
# load data\n",
data_1 = nib.load(nii_path).get_fdata()
# get middle slices\n",
slc_data1 = np.rot90(data_1[np.shape(data_1)[0]//2,:,:])
slc_data2 = np.rot90(data_1[:,np.shape(data_1)[1]//2,:])
slc_data3 = np.rot90(data_1[:,:,np.shape(data_1)[2]//2])
# show slices\n",
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=figsize)
if title: plt.suptitle(title)
axes[0].imshow(slc_data1, cmap=cmap, **imshow_args)
axes[1].imshow(slc_data2, cmap=cmap, **imshow_args)
axes[2].imshow(slc_data3, cmap=cmap, **imshow_args)
axes[0].axis('off')
axes[1].axis('off')
axes[2].axis('off')
fig.tight_layout()
fig.subplots_adjust(top=1.55)
plt.show()
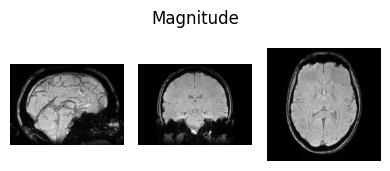
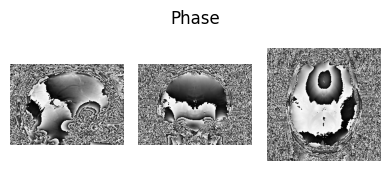
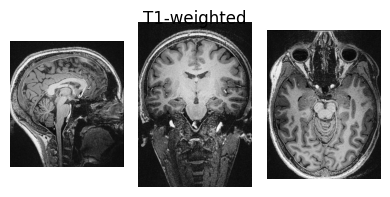
show_nii(glob("bids/sub-*/ses-*/anat/*mag*T2starw*nii*")[0], title="Magnitude", vmax=500)
show_nii(glob("bids/sub-*/ses-*/anat/*phase*T2starw*nii*")[0], title="Phase")
show_nii(glob("bids/sub-*/ses-*/anat/*T1w*nii*")[0], title="T1-weighted")
Interactive Display using Niivue#
from ipyniivue import NiiVue
nv_T1 = NiiVue()
nv_T1.load_volumes([{"path": glob("bids/sub-*/ses-*/anat/*T1w*nii*")[0]}])
nv_T1
[HF-patcher] sub-2: path → url
Run QSMxT#
We are now ready to run QSMxT! We will generate susceptibility maps and segmentations, and export analysis CSVs to file.
The usual way of running QSMxT is to use qsmxt bids_dir. This will launch an interactive command-line interface (CLI) to setup your desired pipelines. However, since we are running this in a notebook, we need to use command-line arguments to by-pass the interface and execute a pipeline.
But first, let’s consider our pipeline settings. For QSM reconstruction, QSMxT provides a range of sensible defaults fit for different purposes. We can list the premade QSM pipelines using --list_premades. For the full pipeline details used for each premade pipeline, see qsm_pipelines.json.
!qsmxt --list_premades
=== Premade pipelines ===
default: Default QSMxT settings (GRE; assumes human brain)
gre: Applies suggested settings for 3D-GRE images
epi: Applies suggested settings for 3D-EPI images (assumes human brain)
bet: Applies a traditional BET-masking approach (artefact reduction unavailable; assumes human brain)
fast: Applies a set of fast algorithms
body: Applies suggested settings for non-brain applications
nextqsm: Applies suggested settings for running the NeXtQSM algorithm (assumes human brain)
[INFO]: Finished
For this demonstration, we will go with the fast pipeline. To export segmentations and analysis results, we will use --do_segmentation and --do_analysis. The --auto_yes option avoid the interactive CLI interface that cannot be used in a notebook:
!qsmxt bids \
--premade fast \
--do_qsm \
--do_segmentation \
--do_analysis \
--auto_yes
[INFO]: QSMxT v8.1.1
[INFO]: Python interpreter: /opt/miniconda-4.12.0/bin/python3.8
[INFO]: Command: qsmxt /home/jovyan/workspace/books/examples/quantitative_imaging/bids --premade 'fast' --do_qsm --do_segmentation --do_analysis --auto_yes
[WARNING]: Pipeline is NOT guidelines compliant (see https://doi.org/10.1002/mrm.30006):; Phase-quality-based masking recommended
[INFO]: Available memory: 94.476 GB
[INFO]: Creating QSMxT workflow for sub-1.ses-20170705.acq-acqmp2ragehighres0p5isoslab.T1w...
[INFO]: Creating QSMxT workflow for sub-1.ses-20170705.acq-acqqsmp21mmisote20.T2starw...
[INFO]: Creating QSMxT workflow for sub-2.ses-20170705.acq-acqmp2ragehighres0p5isoslab.T1w...
[INFO]: Creating QSMxT workflow for sub-2.ses-20170705.acq-acqqsmp21mmisote20.T2starw...
[INFO]: Running using MultiProc plugin with n_procs=32
260409-02:05:21,880 nipype.workflow INFO:
Workflow qsmxt-workflow settings: ['check', 'execution', 'logging', 'monitoring']
260409-02:05:21,943 nipype.workflow INFO:
Running in parallel.
260409-02:05:21,953 nipype.workflow INFO:
[MultiProc] Running 0 tasks, and 24 jobs ready. Free memory (GB): 113.21/113.21, Free processors: 32/32.
260409-02:05:22,23 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-me".
260409-02:05:22,211 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-se".
260409-02:05:22,211 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-me" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/func_read-json-me".
260409-02:05:22,213 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-me".
260409-02:05:22,214 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_read-nii".
260409-02:05:22,216 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-se" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/func_read-json-se".
260409-02:05:22,218 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-se".
260409-02:05:22,218 nipype.workflow INFO:
[Node] Executing "func_read-json-me" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,218 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_read-nii" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/nibabel_read-nii".
260409-02:05:22,220 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_read-nii".
260409-02:05:22,222 nipype.workflow INFO:
[Node] Finished "func_read-json-me", elapsed time 0.001514s.
260409-02:05:22,223 nipype.workflow INFO:
[Node] Executing "func_read-json-se" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,226 nipype.workflow INFO:
[Node] Executing "nibabel_read-nii" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,228 nipype.workflow INFO:
[Node] Finished "func_read-json-se", elapsed time 0.001538s.
260409-02:05:22,237 nipype.workflow INFO:
[Node] Finished "nibabel_read-nii", elapsed time 0.00864s.
260409-02:05:22,217 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_as-canonical".
260409-02:05:22,270 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1".
260409-02:05:22,272 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_as-canonical" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/nibabel_as-canonical".
260409-02:05:22,274 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm".
260409-02:05:22,274 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_as-canonical".
260409-02:05:22,276 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-me".
260409-02:05:22,278 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-se".
260409-02:05:22,279 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/ants_register-t1-to-qsm".
260409-02:05:22,280 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_read-nii".
260409-02:05:22,281 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm".
260409-02:05:22,280 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-me" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/func_read-json-me".
260409-02:05:22,282 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_as-canonical".
260409-02:05:22,282 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-se" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/func_read-json-se".
260409-02:05:22,284 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-se".
260409-02:05:22,284 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1".
260409-02:05:22,285 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_as-canonical" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_as-canonical".
260409-02:05:22,287 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_as-canonical".
260409-02:05:22,288 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/fastsurfer_segment-t1".
260409-02:05:22,289 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1".
260409-02:05:22,292 nipype.workflow INFO:
[Node] Executing "nibabel_as-canonical" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,306 nipype.workflow INFO:
[Node] Executing "fastsurfer_segment-t1" <qsmxt.interfaces.nipype_interface_fastsurfer.FastSurferInterface>
260409-02:05:22,307 nipype.workflow INFO:
[Node] Finished "nibabel_as-canonical", elapsed time 0.011114s.
260409-02:05:22,276 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/fastsurfer_segment-t1".
260409-02:05:22,328 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1".
260409-02:05:22,280 nipype.workflow INFO:
[Node] Executing "nibabel_as-canonical" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,283 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-me".
260409-02:05:22,283 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_read-nii" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_read-nii".
260409-02:05:22,286 nipype.workflow INFO:
[Node] Executing "ants_register-t1-to-qsm" <nipype.interfaces.ants.registration.RegistrationSynQuick>
260409-02:05:22,337 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_read-nii".
260409-02:05:22,289 nipype.workflow INFO:
[Node] Executing "func_read-json-se" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,343 nipype.workflow INFO:
[Node] Finished "nibabel_as-canonical", elapsed time 0.009938s.
260409-02:05:22,344 nipype.workflow INFO:
[Node] Executing "nibabel_read-nii" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,346 nipype.workflow INFO:
[Node] Executing "fastsurfer_segment-t1" <qsmxt.interfaces.nipype_interface_fastsurfer.FastSurferInterface>
260409-02:05:22,356 nipype.workflow INFO:
[Node] Finished "nibabel_read-nii", elapsed time 0.007475s.
260409-02:05:22,286 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-me".
260409-02:05:22,340 nipype.workflow INFO:
[Node] Executing "func_read-json-me" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,392 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-se".
260409-02:05:22,393 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-me" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/func_read-json-me".
260409-02:05:22,395 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-me".
260409-02:05:22,396 nipype.workflow INFO:
[Node] Finished "func_read-json-me", elapsed time 0.001674s.
260409-02:05:22,396 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-se" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/func_read-json-se".
260409-02:05:22,398 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-se".
260409-02:05:22,400 nipype.workflow INFO:
[Node] Executing "func_read-json-me" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,404 nipype.workflow INFO:
[Node] Executing "func_read-json-se" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:22,405 nipype.workflow INFO:
[Node] Finished "func_read-json-me", elapsed time 0.001721s.
260409-02:05:22,408 nipype.workflow INFO:
[Node] Finished "func_read-json-se", elapsed time 0.001373s.
260409-02:05:22,346 nipype.workflow INFO:
[Node] Finished "func_read-json-se", elapsed time 0.00269s.
260409-02:05:23,957 nipype.workflow INFO:
[Job 0] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-me).
260409-02:05:23,959 nipype.workflow INFO:
[Job 1] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-se).
260409-02:05:23,961 nipype.workflow INFO:
[Job 2] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_read-nii).
260409-02:05:23,963 nipype.workflow INFO:
[Job 3] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_as-canonical).
260409-02:05:23,964 nipype.workflow INFO:
[Job 6] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-me).
260409-02:05:23,966 nipype.workflow INFO:
[Job 7] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-se).
260409-02:05:23,967 nipype.workflow INFO:
[Job 8] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_read-nii).
260409-02:05:23,969 nipype.workflow INFO:
[Job 9] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_as-canonical).
260409-02:05:23,971 nipype.workflow INFO:
[Job 12] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-me).
260409-02:05:23,973 nipype.workflow INFO:
[Job 13] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_read-json-se).
260409-02:05:23,976 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 14 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:05:24,90 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm".
260409-02:05:24,94 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_read-nii".
260409-02:05:24,96 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_as-canonical".
260409-02:05:24,97 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/ants_register-t1-to-qsm".
260409-02:05:24,98 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_read-nii" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/nibabel_read-nii".
260409-02:05:24,99 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-me".
260409-02:05:24,100 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_read-nii".
260409-02:05:24,100 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-se".
260409-02:05:24,100 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_as-canonical" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/nibabel_as-canonical".
260409-02:05:24,103 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_as-canonical".
260409-02:05:24,105 nipype.workflow INFO:
[Node] Executing "nibabel_read-nii" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:24,109 nipype.workflow INFO:
[Node] Executing "nibabel_as-canonical" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:24,114 nipype.workflow INFO:
[Node] Finished "nibabel_read-nii", elapsed time 0.006835s.
260409-02:05:24,115 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-se" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/func_read-json-se".
260409-02:05:24,115 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-me" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/func_read-json-me".
260409-02:05:24,117 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-me".
260409-02:05:24,121 nipype.workflow INFO:
[Node] Finished "nibabel_as-canonical", elapsed time 0.009549s.
260409-02:05:24,123 nipype.workflow INFO:
[Node] Executing "func_read-json-me" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:24,128 nipype.workflow INFO:
[Node] Finished "func_read-json-me", elapsed time 0.001781s.
260409-02:05:24,98 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm".
260409-02:05:24,156 nipype.workflow INFO:
[Node] Executing "ants_register-t1-to-qsm" <nipype.interfaces.ants.registration.RegistrationSynQuick>
260409-02:05:24,116 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-se".
260409-02:05:24,172 nipype.workflow INFO:
[Node] Executing "func_read-json-se" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:24,175 nipype.workflow INFO:
[Node] Finished "func_read-json-se", elapsed time 0.001304s.
260409-02:05:25,957 nipype.workflow INFO:
[Job 14] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_read-nii).
260409-02:05:25,959 nipype.workflow INFO:
[Job 15] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_as-canonical).
260409-02:05:25,960 nipype.workflow INFO:
[Job 18] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-me).
260409-02:05:25,962 nipype.workflow INFO:
[Job 19] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_read-json-se).
260409-02:05:25,965 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 10 jobs ready. Free memory (GB): 73.21/113.21, Free processors: 4/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:05:26,48 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_read-nii".
260409-02:05:26,51 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_as-canonical".
260409-02:05:26,52 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_read-nii" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_read-nii".
260409-02:05:26,54 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_read-nii".
260409-02:05:26,55 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_getfirst-canonical-magnitude".
260409-02:05:26,55 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_as-canonical" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_as-canonical".
260409-02:05:26,58 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_as-canonical".
260409-02:05:26,58 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_scale-phase".
260409-02:05:26,59 nipype.workflow INFO:
[Node] Executing "nibabel_read-nii" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:26,62 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_scale-phase" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_scale-phase".
260409-02:05:26,63 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_scale-phase".
260409-02:05:26,66 nipype.workflow INFO:
[Node] Finished "nibabel_read-nii", elapsed time 0.004186s.
260409-02:05:26,69 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_scale-phase" <qsmxt.interfaces.nipype_interface_processphase.ScalePhaseInterface>
260409-02:05:26,59 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_getfirst-canonical-magnitude" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/func_getfirst-canonical-magnitude".
260409-02:05:26,112 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_getfirst-canonical-magnitude".
260409-02:05:26,63 nipype.workflow INFO:
[Node] Executing "nibabel_as-canonical" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:26,119 nipype.workflow INFO:
[Node] Executing "func_getfirst-canonical-magnitude" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:26,124 nipype.workflow INFO:
[Node] Finished "func_getfirst-canonical-magnitude", elapsed time 0.001424s.
260409-02:05:26,132 nipype.workflow INFO:
[Node] Finished "nibabel_as-canonical", elapsed time 0.014026s.
260409-02:05:26,435 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_scale-phase", elapsed time 0.363988s.
260409-02:05:27,970 nipype.workflow INFO:
[Job 20] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_read-nii).
260409-02:05:27,972 nipype.workflow INFO:
[Job 21] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_as-canonical).
260409-02:05:27,982 nipype.workflow INFO:
[Job 24] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_getfirst-canonical-magnitude).
260409-02:05:27,983 nipype.workflow INFO:
[Job 26] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_scale-phase).
260409-02:05:27,985 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 9 jobs ready. Free memory (GB): 73.21/113.21, Free processors: 4/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:05:28,68 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_getfirst-canonical-magnitude".
260409-02:05:28,72 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_getfirst-canonical-magnitude".
260409-02:05:28,72 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_getfirst-canonical-magnitude" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/func_getfirst-canonical-magnitude".
260409-02:05:28,74 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_getfirst-canonical-magnitude".
260409-02:05:28,74 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_scale-phase".
260409-02:05:28,76 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_getfirst-canonical-magnitude" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/func_getfirst-canonical-magnitude".
260409-02:05:28,77 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_getfirst-canonical-magnitude".
260409-02:05:28,78 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_scale-phase" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_scale-phase".
260409-02:05:28,79 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_scale-phase".
260409-02:05:28,81 nipype.workflow INFO:
[Node] Executing "func_getfirst-canonical-magnitude" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:28,83 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_scale-phase" <qsmxt.interfaces.nipype_interface_processphase.ScalePhaseInterface>
260409-02:05:28,84 nipype.workflow INFO:
[Node] Finished "func_getfirst-canonical-magnitude", elapsed time 0.00099s.
260409-02:05:28,77 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_getfirst-canonical-magnitude".
260409-02:05:28,78 nipype.workflow INFO:
[Node] Executing "func_getfirst-canonical-magnitude" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:28,131 nipype.workflow INFO:
[Node] Finished "func_getfirst-canonical-magnitude", elapsed time 0.000867s.
260409-02:05:28,132 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_getfirst-canonical-magnitude" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/func_getfirst-canonical-magnitude".
260409-02:05:28,134 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_getfirst-canonical-magnitude".
260409-02:05:28,138 nipype.workflow INFO:
[Node] Executing "func_getfirst-canonical-magnitude" <nipype.interfaces.utility.wrappers.Function>
260409-02:05:28,141 nipype.workflow INFO:
[Node] Finished "func_getfirst-canonical-magnitude", elapsed time 0.000988s.
260409-02:05:28,439 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_scale-phase", elapsed time 0.354448s.
260409-02:05:29,961 nipype.workflow INFO:
[Job 27] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_getfirst-canonical-magnitude).
260409-02:05:29,962 nipype.workflow INFO:
[Job 29] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.func_getfirst-canonical-magnitude).
260409-02:05:29,964 nipype.workflow INFO:
[Job 31] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_scale-phase).
260409-02:05:29,965 nipype.workflow INFO:
[Job 32] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.func_getfirst-canonical-magnitude).
260409-02:05:29,968 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 6 jobs ready. Free memory (GB): 73.21/113.21, Free processors: 4/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:05:30,42 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_axial-resampling".
260409-02:05:30,47 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_axial-resampling".
260409-02:05:30,51 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_axial-resampling" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_nilearn_axial-resampling".
260409-02:05:30,53 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_axial-resampling".
260409-02:05:30,58 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_nilearn_axial-resampling" <qsmxt.interfaces.nipype_interface_axialsampling.AxialSamplingInterface>
260409-02:05:30,65 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_nilearn_axial-resampling", elapsed time 0.005727s.
260409-02:05:30,47 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_axial-resampling" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_nilearn_axial-resampling".
260409-02:05:30,99 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_axial-resampling".
260409-02:05:30,103 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_nilearn_axial-resampling" <qsmxt.interfaces.nipype_interface_axialsampling.AxialSamplingInterface>
260409-02:05:30,111 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_nilearn_axial-resampling", elapsed time 0.005214s.
260409-02:05:31,962 nipype.workflow INFO:
[Job 36] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_axial-resampling).
260409-02:05:31,963 nipype.workflow INFO:
[Job 41] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_axial-resampling).
260409-02:05:31,964 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 8 jobs ready. Free memory (GB): 73.21/113.21, Free processors: 4/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:05:32,31 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo".
260409-02:05:32,33 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo".
260409-02:05:32,44 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/qsm_workflow/mrt_romeo".
260409-02:05:32,54 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo".
260409-02:05:32,40 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/qsm_workflow/mrt_romeo".
260409-02:05:32,64 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo".
260409-02:05:32,88 nipype.workflow INFO:
[Node] Executing "mrt_romeo" <qsmxt.interfaces.nipype_interface_romeo.RomeoB0Interface>
260409-02:05:32,88 nipype.workflow INFO:
[Node] Executing "mrt_romeo" <qsmxt.interfaces.nipype_interface_romeo.RomeoB0Interface>
260409-02:05:33,966 nipype.workflow INFO:
[MultiProc] Running 6 tasks, and 6 jobs ready. Free memory (GB): 68.53/113.21, Free processors: 2/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:06:18,952 nipype.workflow INFO:
[Node] Finished "mrt_romeo", elapsed time 46.861514s.
260409-02:06:20,10 nipype.workflow INFO:
[Job 50] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo).
260409-02:06:20,12 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 7 jobs ready. Free memory (GB): 70.87/113.21, Free processors: 3/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:06:20,115 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.nibabel-numpy_normalize-phase".
260409-02:06:20,119 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.nibabel-numpy_normalize-phase" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/qsm_workflow/nibabel-numpy_normalize-phase".
260409-02:06:20,122 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.nibabel-numpy_normalize-phase".
260409-02:06:20,144 nipype.workflow INFO:
[Node] Executing "nibabel-numpy_normalize-phase" <qsmxt.interfaces.nipype_interface_processphase.PhaseToNormalizedInterface>
260409-02:06:20,351 nipype.workflow INFO:
[Node] Finished "nibabel-numpy_normalize-phase", elapsed time 0.203485s.
260409-02:06:20,787 nipype.workflow INFO:
[Node] Finished "mrt_romeo", elapsed time 48.695164s.
260409-02:06:22,13 nipype.workflow INFO:
[Job 46] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.mrt_romeo).
260409-02:06:22,15 nipype.workflow INFO:
[Job 55] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.nibabel-numpy_normalize-phase).
260409-02:06:22,18 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 7 jobs ready. Free memory (GB): 73.21/113.21, Free processors: 4/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:06:22,107 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.nibabel-numpy_normalize-phase".
260409-02:06:22,110 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.nibabel-numpy_normalize-phase" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/qsm_workflow/nibabel-numpy_normalize-phase".
260409-02:06:22,111 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.nibabel-numpy_normalize-phase".
260409-02:06:22,123 nipype.workflow INFO:
[Node] Executing "nibabel-numpy_normalize-phase" <qsmxt.interfaces.nipype_interface_processphase.PhaseToNormalizedInterface>
260409-02:06:22,234 nipype.workflow INFO:
[Node] Finished "nibabel-numpy_normalize-phase", elapsed time 0.109236s.
260409-02:06:24,15 nipype.workflow INFO:
[Job 53] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.nibabel-numpy_normalize-phase).
260409-02:06:24,18 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 6 jobs ready. Free memory (GB): 73.21/113.21, Free processors: 4/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:08:07,947 nipype.workflow INFO:
[Node] Finished "ants_register-t1-to-qsm", elapsed time 163.788923s.
260409-02:08:08,130 nipype.workflow INFO:
[Job 11] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm).
260409-02:08:08,132 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 6 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:08:08,261 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1".
260409-02:08:08,265 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/fastsurfer_segment-t1".
260409-02:08:08,266 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1".
260409-02:08:08,288 nipype.workflow INFO:
[Node] Executing "fastsurfer_segment-t1" <qsmxt.interfaces.nipype_interface_fastsurfer.FastSurferInterface>
260409-02:08:10,133 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 5 jobs ready. Free memory (GB): 69.21/113.21, Free processors: 2/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:10:19,930 nipype.workflow INFO:
[Node] Finished "ants_register-t1-to-qsm", elapsed time 297.590371s.
260409-02:10:22,336 nipype.workflow INFO:
[Job 5] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm).
260409-02:10:22,338 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 5 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:10:22,534 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm".
260409-02:10:22,549 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/ants_register-t1-to-qsm".
260409-02:10:22,550 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm".
260409-02:10:22,554 nipype.workflow INFO:
[Node] Executing "ants_register-t1-to-qsm" <nipype.interfaces.ants.registration.RegistrationSynQuick>
260409-02:10:24,347 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 4 jobs ready. Free memory (GB): 69.21/113.21, Free processors: 2/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:16:54,152 nipype.workflow INFO:
[Node] Finished "ants_register-t1-to-qsm", elapsed time 391.596412s.
260409-02:16:54,932 nipype.workflow INFO:
[Job 17] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_register-t1-to-qsm).
260409-02:16:54,935 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 4 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:16:55,47 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1".
260409-02:16:55,51 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/fastsurfer_segment-t1".
260409-02:16:55,51 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1".
260409-02:16:55,65 nipype.workflow INFO:
[Node] Executing "fastsurfer_segment-t1" <qsmxt.interfaces.nipype_interface_fastsurfer.FastSurferInterface>
260409-02:16:56,935 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 3 jobs ready. Free memory (GB): 65.21/113.21, Free processors: 0/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:29,625 nipype.workflow INFO:
[Node] Finished "fastsurfer_segment-t1", elapsed time 727.275809s.
260409-02:17:30,998 nipype.workflow INFO:
[Job 4] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1).
260409-02:17:31,0 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 4 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
260409-02:17:31,69 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm".
260409-02:17:31,86 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.numpy_numpy_nibabel_mgz2nii".
260409-02:17:31,87 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/ants_register-t1-to-qsm".
260409-02:17:31,88 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm".
260409-02:17:31,88 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.numpy_numpy_nibabel_mgz2nii" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/numpy_numpy_nibabel_mgz2nii".
260409-02:17:31,89 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.numpy_numpy_nibabel_mgz2nii".
260409-02:17:31,91 nipype.workflow INFO:
[Node] Executing "ants_register-t1-to-qsm" <nipype.interfaces.ants.registration.RegistrationSynQuick>
260409-02:17:31,92 nipype.workflow INFO:
[Node] Executing "numpy_numpy_nibabel_mgz2nii" <qsmxt.interfaces.nipype_interface_mgz2nii.Mgz2NiiInterface>
260409-02:17:31,362 nipype.workflow INFO:
[Node] Finished "numpy_numpy_nibabel_mgz2nii", elapsed time 0.267635s.
260409-02:17:32,545 nipype.workflow INFO:
[Node] Finished "fastsurfer_segment-t1", elapsed time 730.236948s.
260409-02:17:33,2 nipype.workflow INFO:
[Job 10] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1).
260409-02:17:33,4 nipype.workflow INFO:
[Job 25] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.numpy_numpy_nibabel_mgz2nii).
260409-02:17:33,5 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 5 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:33,112 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.numpy_numpy_nibabel_mgz2nii".
260409-02:17:33,115 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.numpy_numpy_nibabel_mgz2nii" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/numpy_numpy_nibabel_mgz2nii".
260409-02:17:33,116 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.numpy_numpy_nibabel_mgz2nii".
260409-02:17:33,118 nipype.workflow INFO:
[Node] Executing "numpy_numpy_nibabel_mgz2nii" <qsmxt.interfaces.nipype_interface_mgz2nii.Mgz2NiiInterface>
260409-02:17:33,116 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:17:33,171 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:17:33,172 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:17:33,175 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm".
260409-02:17:33,183 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/ants_transform-segmentation-to-qsm".
260409-02:17:33,185 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm".
260409-02:17:33,200 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_nilearn_t1w-seg-resampled" <qsmxt.interfaces.nipype_interface_resample_like.ResampleLikeInterface>
260409-02:17:33,352 nipype.workflow INFO:
[Node] Executing "ants_transform-segmentation-to-qsm" <nipype.interfaces.ants.resampling.ApplyTransforms>
260409-02:17:33,386 nipype.workflow INFO:
[Node] Finished "numpy_numpy_nibabel_mgz2nii", elapsed time 0.266793s.
260409-02:17:35,4 nipype.workflow INFO:
[Job 28] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.numpy_numpy_nibabel_mgz2nii).
260409-02:17:35,7 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 4 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:35,210 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:17:35,214 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:17:35,215 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:17:35,228 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_transform-segmentation-to-qsm".
260409-02:17:35,234 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_transform-segmentation-to-qsm" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/ants_transform-segmentation-to-qsm".
260409-02:17:35,235 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_transform-segmentation-to-qsm".
260409-02:17:35,248 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_nilearn_t1w-seg-resampled" <qsmxt.interfaces.nipype_interface_resample_like.ResampleLikeInterface>
260409-02:17:35,284 nipype.workflow INFO:
[Node] Executing "ants_transform-segmentation-to-qsm" <nipype.interfaces.ants.resampling.ApplyTransforms>
260409-02:17:35,765 nipype.workflow INFO:
[Node] Finished "ants_transform-segmentation-to-qsm", elapsed time 2.410821s.
260409-02:17:36,629 nipype.workflow INFO:
[Node] Finished "ants_transform-segmentation-to-qsm", elapsed time 1.3151570000000001s.
260409-02:17:37,6 nipype.workflow INFO:
[Job 35] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm).
260409-02:17:37,7 nipype.workflow INFO:
[Job 38] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_transform-segmentation-to-qsm).
260409-02:17:37,8 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 3 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:37,111 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet".
260409-02:17:37,115 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/mask_workflow/fsl-bet".
260409-02:17:37,116 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet".
260409-02:17:37,120 nipype.workflow INFO:
[Node] Executing "fsl-bet" <qsmxt.interfaces.nipype_interface_bet2.Bet2Interface>
260409-02:17:39,10 nipype.workflow INFO:
[MultiProc] Running 6 tasks, and 2 jobs ready. Free memory (GB): 75.21/113.21, Free processors: 0/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:40,690 nipype.workflow INFO:
[Node] Finished "fsl-bet", elapsed time 3.569454s.
260409-02:17:41,10 nipype.workflow INFO:
[Job 45] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet).
260409-02:17:41,13 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 3 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:41,83 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists2".
260409-02:17:41,86 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.scipy_numpy_nibabel_bet_erode".
260409-02:17:41,86 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists2" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/combine_lists2".
260409-02:17:41,87 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists2".
260409-02:17:41,90 nipype.workflow INFO:
[Node] Executing "combine_lists2" <nipype.interfaces.utility.wrappers.Function>
260409-02:17:41,89 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.scipy_numpy_nibabel_bet_erode" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/mask_workflow/scipy_numpy_nibabel_bet_erode".
260409-02:17:41,90 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.scipy_numpy_nibabel_bet_erode".
260409-02:17:41,91 nipype.workflow INFO:
[Node] Finished "combine_lists2", elapsed time 0.000662s.
260409-02:17:41,95 nipype.workflow INFO:
[Node] Executing "scipy_numpy_nibabel_bet_erode" <qsmxt.interfaces.nipype_interface_erode.ErosionInterface>
260409-02:17:41,988 nipype.workflow INFO:
[Node] Finished "scipy_numpy_nibabel_bet_erode", elapsed time 0.891349s.
260409-02:17:43,12 nipype.workflow INFO:
[Job 47] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists2).
260409-02:17:43,14 nipype.workflow INFO:
[Job 52] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.scipy_numpy_nibabel_bet_erode).
260409-02:17:43,15 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 2 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:43,147 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet".
260409-02:17:43,150 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/mask_workflow/fsl-bet".
260409-02:17:43,151 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet".
260409-02:17:43,165 nipype.workflow INFO:
[Node] Executing "fsl-bet" <qsmxt.interfaces.nipype_interface_bet2.Bet2Interface>
260409-02:17:44,965 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_nilearn_t1w-seg-resampled", elapsed time 11.763157s.
260409-02:17:45,15 nipype.workflow INFO:
[Job 34] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled).
260409-02:17:45,17 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 2 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 1/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:45,113 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.copyfile".
260409-02:17:45,138 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.copyfile" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/copyfile".
260409-02:17:45,139 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.copyfile".
260409-02:17:45,149 nipype.workflow INFO:
[Node] Executing "copyfile" <qsmxt.interfaces.nipype_interface_copyfile.DynamicCopyFiles>
260409-02:17:45,411 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_nilearn_t1w-seg-resampled", elapsed time 10.162106s.
260409-02:17:46,25 nipype.workflow INFO:
[Node] Finished "copyfile", elapsed time 0.874857s.
260409-02:17:46,999 nipype.workflow INFO:
[Node] Finished "fsl-bet", elapsed time 3.83186s.
260409-02:17:47,17 nipype.workflow INFO:
[Job 37] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled).
260409-02:17:47,18 nipype.workflow INFO:
[Job 49] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.fsl-bet).
260409-02:17:47,19 nipype.workflow INFO:
[Job 44] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.copyfile).
260409-02:17:47,20 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 2 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:47,109 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.scipy_numpy_nibabel_bet_erode".
260409-02:17:47,113 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp".
260409-02:17:47,113 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.scipy_numpy_nibabel_bet_erode" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/mask_workflow/scipy_numpy_nibabel_bet_erode".
260409-02:17:47,114 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.scipy_numpy_nibabel_bet_erode".
260409-02:17:47,118 nipype.workflow INFO:
[Node] Executing "scipy_numpy_nibabel_bet_erode" <qsmxt.interfaces.nipype_interface_erode.ErosionInterface>
260409-02:17:47,120 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/qsm_workflow/qsmjl_vsharp".
260409-02:17:47,121 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp".
260409-02:17:47,157 nipype.workflow INFO:
[Node] Executing "qsmjl_vsharp" <qsmxt.interfaces.nipype_interface_qsmjl.VsharpInterface>
260409-02:17:48,86 nipype.workflow INFO:
[Node] Finished "scipy_numpy_nibabel_bet_erode", elapsed time 0.967154s.
260409-02:17:49,18 nipype.workflow INFO:
[Job 54] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.mask_workflow.scipy_numpy_nibabel_bet_erode).
260409-02:17:49,20 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 1 jobs ready. Free memory (GB): 79.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:17:49,117 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp".
260409-02:17:49,121 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/qsm_workflow/qsmjl_vsharp".
260409-02:17:49,122 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp".
260409-02:17:49,153 nipype.workflow INFO:
[Node] Executing "qsmjl_vsharp" <qsmxt.interfaces.nipype_interface_qsmjl.VsharpInterface>
260409-02:17:51,21 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 0 jobs ready. Free memory (GB): 77.21/113.21, Free processors: 6/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:18:13,549 nipype.workflow INFO:
[Node] Finished "qsmjl_vsharp", elapsed time 26.39032s.
260409-02:18:15,48 nipype.workflow INFO:
[Job 56] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp).
260409-02:18:15,51 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 1 jobs ready. Free memory (GB): 79.21/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:18:15,163 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts".
260409-02:18:15,168 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/qsm_workflow/qsmjl_rts".
260409-02:18:15,170 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts".
260409-02:18:15,200 nipype.workflow INFO:
[Node] Executing "qsmjl_rts" <qsmxt.interfaces.nipype_interface_qsmjl.RtsQsmInterface>
260409-02:18:16,350 nipype.workflow INFO:
[Node] Finished "qsmjl_vsharp", elapsed time 27.195776s.
260409-02:18:17,49 nipype.workflow INFO:
[Job 57] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_vsharp).
260409-02:18:17,51 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 1 jobs ready. Free memory (GB): 78.80/113.21, Free processors: 8/32.
Currently running:
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:18:17,210 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts".
260409-02:18:17,215 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/qsm_workflow/qsmjl_rts".
260409-02:18:17,216 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts".
260409-02:18:17,238 nipype.workflow INFO:
[Node] Executing "qsmjl_rts" <qsmxt.interfaces.nipype_interface_qsmjl.RtsQsmInterface>
260409-02:18:19,51 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 0 jobs ready. Free memory (GB): 76.39/113.21, Free processors: 6/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts
* qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:18:51,195 nipype.workflow INFO:
[Node] Finished "qsmjl_rts", elapsed time 33.955308s.
260409-02:18:51,983 nipype.workflow INFO:
[Node] Finished "qsmjl_rts", elapsed time 36.781201s.
260409-02:18:53,89 nipype.workflow INFO:
[Job 58] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts).
260409-02:18:53,92 nipype.workflow INFO:
[Job 59] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.qsm_workflow.qsmjl_rts).
260409-02:18:53,97 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 2 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:18:53,210 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-average".
260409-02:18:53,214 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-average".
260409-02:18:53,216 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-average" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_qsm-average".
260409-02:18:53,219 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-average".
260409-02:18:53,223 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_qsm-average" <qsmxt.interfaces.nipype_interface_nonzeroaverage.NonzeroAverageInterface>
260409-02:18:53,230 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_qsm-average", elapsed time 0.000528s.
260409-02:18:53,217 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-average" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_qsm-average".
260409-02:18:53,270 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-average".
260409-02:18:53,273 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_qsm-average" <qsmxt.interfaces.nipype_interface_nonzeroaverage.NonzeroAverageInterface>
260409-02:18:53,275 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_qsm-average", elapsed time 0.000413s.
260409-02:18:55,90 nipype.workflow INFO:
[Job 60] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-average).
260409-02:18:55,92 nipype.workflow INFO:
[Job 61] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-average).
260409-02:18:55,93 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 2 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:18:55,225 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_qsm-resampled".
260409-02:18:55,228 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_qsm-resampled".
260409-02:18:55,233 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_qsm-resampled" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_nilearn_qsm-resampled".
260409-02:18:55,235 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_qsm-resampled" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_nilearn_qsm-resampled".
260409-02:18:55,236 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_qsm-resampled".
260409-02:18:55,239 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_nilearn_qsm-resampled" <qsmxt.interfaces.nipype_interface_resample_like.ResampleLikeInterface>
260409-02:18:55,244 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_nilearn_qsm-resampled", elapsed time 0.0031s.
260409-02:18:55,235 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_qsm-resampled".
260409-02:18:55,291 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_nilearn_qsm-resampled" <qsmxt.interfaces.nipype_interface_resample_like.ResampleLikeInterface>
260409-02:18:55,297 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_nilearn_qsm-resampled", elapsed time 0.004299s.
260409-02:18:57,93 nipype.workflow INFO:
[Job 62] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_qsm-resampled).
260409-02:18:57,94 nipype.workflow INFO:
[Job 63] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_qsm-resampled).
260409-02:18:57,95 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 4 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:18:57,172 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-referenced".
260409-02:18:57,175 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists1".
260409-02:18:57,177 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-referenced".
260409-02:18:57,179 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-referenced" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_qsm-referenced".
260409-02:18:57,180 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-referenced".
260409-02:18:57,179 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/combine_lists1".
260409-02:18:57,181 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists1".
260409-02:18:57,183 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-referenced" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_qsm-referenced".
260409-02:18:57,185 nipype.workflow INFO:
[Node] Executing "combine_lists1" <nipype.interfaces.utility.wrappers.Function>
260409-02:18:57,188 nipype.workflow INFO:
[Node] Finished "combine_lists1", elapsed time 0.001331s.
260409-02:18:57,205 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-referenced".
260409-02:18:57,226 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_qsm-referenced" <qsmxt.interfaces.nipype_interface_qsm_referencing.ReferenceQSMInterface>
260409-02:18:57,227 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_qsm-referenced" <qsmxt.interfaces.nipype_interface_qsm_referencing.ReferenceQSMInterface>
260409-02:18:57,179 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists1".
260409-02:18:57,233 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/combine_lists1".
260409-02:18:57,235 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists1".
260409-02:18:57,240 nipype.workflow INFO:
[Node] Executing "combine_lists1" <nipype.interfaces.utility.wrappers.Function>
260409-02:18:57,245 nipype.workflow INFO:
[Node] Finished "combine_lists1", elapsed time 0.003334s.
260409-02:18:57,734 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_qsm-referenced", elapsed time 0.504299s.
260409-02:18:57,869 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_qsm-referenced", elapsed time 0.640778s.
260409-02:18:59,94 nipype.workflow INFO:
[Job 64] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-referenced).
260409-02:18:59,95 nipype.workflow INFO:
[Job 65] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists1).
260409-02:18:59,96 nipype.workflow INFO:
[Job 66] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_qsm-referenced).
260409-02:18:59,96 nipype.workflow INFO:
[Job 67] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists1).
260409-02:18:59,98 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 3 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:18:59,166 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copy_qsm_json_sidecar".
260409-02:18:59,169 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.create_permutations".
260409-02:18:59,171 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copy_qsm_json_sidecar" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/copy_qsm_json_sidecar".
260409-02:18:59,172 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.create_permutations" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/create_permutations".
260409-02:18:59,173 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.create_permutations".
260409-02:18:59,175 nipype.workflow INFO:
[Node] Executing "create_permutations" <nipype.interfaces.utility.wrappers.Function>
260409-02:18:59,177 nipype.workflow INFO:
[Node] Finished "create_permutations", elapsed time 0.000748s.
260409-02:18:59,180 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copy_qsm_json_sidecar".
260409-02:18:59,182 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copy_qsm_json_sidecar" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/copy_qsm_json_sidecar".
260409-02:18:59,183 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copy_qsm_json_sidecar".
260409-02:18:59,185 nipype.workflow INFO:
[Node] Executing "copy_qsm_json_sidecar" <qsmxt.interfaces.nipype_interface_copy_json_sidecar.CopyJsonSidecarInterface>
260409-02:18:59,186 nipype.workflow INFO:
[Node] Finished "copy_qsm_json_sidecar", elapsed time 0.000583s.
260409-02:18:59,172 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copy_qsm_json_sidecar".
260409-02:18:59,227 nipype.workflow INFO:
[Node] Executing "copy_qsm_json_sidecar" <qsmxt.interfaces.nipype_interface_copy_json_sidecar.CopyJsonSidecarInterface>
260409-02:18:59,238 nipype.workflow INFO:
[Node] Finished "copy_qsm_json_sidecar", elapsed time 0.000711s.
260409-02:19:01,97 nipype.workflow INFO:
[Job 68] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copy_qsm_json_sidecar).
260409-02:19:01,99 nipype.workflow INFO:
[Job 69] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.create_permutations).
260409-02:19:01,101 nipype.workflow INFO:
[Job 70] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copy_qsm_json_sidecar).
260409-02:19:01,103 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 1 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:19:03,99 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 4 jobs ready. Free memory (GB): 81.21/113.21, Free processors: 10/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:19:03,192 nipype.workflow INFO:
[Node] Outdated cache found for "_nibabel_numpy_analyse-qsm0".
260409-02:19:03,197 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm0" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm0".
260409-02:19:03,198 nipype.workflow INFO:
[Node] Outdated cache found for "_nibabel_numpy_analyse-qsm0".
260409-02:19:03,201 nipype.workflow INFO:
[Node] Executing "_nibabel_numpy_analyse-qsm0" <qsmxt.interfaces.nipype_interface_analyse.AnalyseInterface>
260409-02:19:03,205 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm1".
260409-02:19:03,206 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm2" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm2".
260409-02:19:03,206 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm3" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm3".
260409-02:19:03,208 nipype.workflow INFO:
[Node] Executing "_nibabel_numpy_analyse-qsm1" <qsmxt.interfaces.nipype_interface_analyse.AnalyseInterface>
260409-02:19:03,210 nipype.workflow INFO:
[Node] Executing "_nibabel_numpy_analyse-qsm3" <qsmxt.interfaces.nipype_interface_analyse.AnalyseInterface>
260409-02:19:03,226 nipype.workflow INFO:
[Node] Executing "_nibabel_numpy_analyse-qsm2" <qsmxt.interfaces.nipype_interface_analyse.AnalyseInterface>
260409-02:19:05,101 nipype.workflow INFO:
[MultiProc] Running 7 tasks, and 0 jobs ready. Free memory (GB): 73.21/113.21, Free processors: 6/32.
Currently running:
* _nibabel_numpy_analyse-qsm3
* _nibabel_numpy_analyse-qsm2
* _nibabel_numpy_analyse-qsm1
* _nibabel_numpy_analyse-qsm0
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:20:00,718 nipype.workflow INFO:
[Node] Finished "_nibabel_numpy_analyse-qsm2", elapsed time 57.491037s.
260409-02:20:00,890 nipype.workflow INFO:
[Node] Finished "ants_register-t1-to-qsm", elapsed time 149.797591s.
260409-02:20:01,154 nipype.workflow INFO:
[Job 23] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_register-t1-to-qsm).
260409-02:20:01,155 nipype.workflow INFO:
[Job 78] Completed (_nibabel_numpy_analyse-qsm2).
260409-02:20:01,157 nipype.workflow INFO:
[MultiProc] Running 5 tasks, and 0 jobs ready. Free memory (GB): 83.21/113.21, Free processors: 13/32.
Currently running:
* _nibabel_numpy_analyse-qsm3
* _nibabel_numpy_analyse-qsm1
* _nibabel_numpy_analyse-qsm0
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:20:04,152 nipype.workflow INFO:
[Node] Finished "_nibabel_numpy_analyse-qsm3", elapsed time 60.941507s.
260409-02:20:04,499 nipype.workflow INFO:
[Node] Finished "_nibabel_numpy_analyse-qsm0", elapsed time 61.296299s.
260409-02:20:05,159 nipype.workflow INFO:
[Job 76] Completed (_nibabel_numpy_analyse-qsm0).
260409-02:20:05,161 nipype.workflow INFO:
[Job 79] Completed (_nibabel_numpy_analyse-qsm3).
260409-02:20:05,163 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 0 jobs ready. Free memory (GB): 87.21/113.21, Free processors: 15/32.
Currently running:
* _nibabel_numpy_analyse-qsm1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:20:05,280 nipype.workflow INFO:
[Node] Finished "_nibabel_numpy_analyse-qsm1", elapsed time 62.071232s.
260409-02:20:07,161 nipype.workflow INFO:
[Job 77] Completed (_nibabel_numpy_analyse-qsm1).
260409-02:20:07,163 nipype.workflow INFO:
[MultiProc] Running 2 tasks, and 1 jobs ready. Free memory (GB): 89.21/113.21, Free processors: 16/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:20:07,262 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_analyse-qsm".
260409-02:20:07,267 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_analyse-qsm".
260409-02:20:07,272 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm0" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm0".
260409-02:20:07,274 nipype.workflow INFO:
[Node] Cached "_nibabel_numpy_analyse-qsm0" - collecting precomputed outputs
260409-02:20:07,274 nipype.workflow INFO:
[Node] "_nibabel_numpy_analyse-qsm0" found cached.
260409-02:20:07,275 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm1".
260409-02:20:07,276 nipype.workflow INFO:
[Node] Cached "_nibabel_numpy_analyse-qsm1" - collecting precomputed outputs
260409-02:20:07,276 nipype.workflow INFO:
[Node] "_nibabel_numpy_analyse-qsm1" found cached.
260409-02:20:07,277 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm2" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm2".
260409-02:20:07,278 nipype.workflow INFO:
[Node] Cached "_nibabel_numpy_analyse-qsm2" - collecting precomputed outputs
260409-02:20:07,279 nipype.workflow INFO:
[Node] "_nibabel_numpy_analyse-qsm2" found cached.
260409-02:20:07,280 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm3" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm3".
260409-02:20:07,281 nipype.workflow INFO:
[Node] Cached "_nibabel_numpy_analyse-qsm3" - collecting precomputed outputs
260409-02:20:07,281 nipype.workflow INFO:
[Node] "_nibabel_numpy_analyse-qsm3" found cached.
260409-02:20:09,166 nipype.workflow INFO:
[Job 72] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_analyse-qsm).
260409-02:20:09,169 nipype.workflow INFO:
[MultiProc] Running 2 tasks, and 1 jobs ready. Free memory (GB): 89.21/113.21, Free processors: 16/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:20:09,319 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copyfile".
260409-02:20:09,324 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copyfile" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-1/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/copyfile".
260409-02:20:09,325 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copyfile".
260409-02:20:09,330 nipype.workflow INFO:
[Node] Executing "copyfile" <qsmxt.interfaces.nipype_interface_copyfile.DynamicCopyFiles>
260409-02:20:09,563 nipype.workflow INFO:
[Node] Finished "copyfile", elapsed time 0.231104s.
260409-02:20:11,171 nipype.workflow INFO:
[Job 74] Completed (qsmxt-workflow.sub-1.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copyfile).
260409-02:20:11,174 nipype.workflow INFO:
[MultiProc] Running 2 tasks, and 0 jobs ready. Free memory (GB): 89.21/113.21, Free processors: 16/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1
260409-02:20:11,773 nipype.workflow INFO:
[Node] Finished "fastsurfer_segment-t1", elapsed time 723.48346s.
260409-02:20:13,173 nipype.workflow INFO:
[Job 16] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.fastsurfer_segment-t1).
260409-02:20:13,175 nipype.workflow INFO:
[MultiProc] Running 1 tasks, and 1 jobs ready. Free memory (GB): 101.21/113.21, Free processors: 24/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
260409-02:20:13,255 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.numpy_numpy_nibabel_mgz2nii".
260409-02:20:13,257 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.numpy_numpy_nibabel_mgz2nii" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/numpy_numpy_nibabel_mgz2nii".
260409-02:20:13,258 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.numpy_numpy_nibabel_mgz2nii".
260409-02:20:13,260 nipype.workflow INFO:
[Node] Executing "numpy_numpy_nibabel_mgz2nii" <qsmxt.interfaces.nipype_interface_mgz2nii.Mgz2NiiInterface>
260409-02:20:13,471 nipype.workflow INFO:
[Node] Finished "numpy_numpy_nibabel_mgz2nii", elapsed time 0.209968s.
260409-02:20:15,175 nipype.workflow INFO:
[Job 30] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.numpy_numpy_nibabel_mgz2nii).
260409-02:20:15,178 nipype.workflow INFO:
[MultiProc] Running 1 tasks, and 2 jobs ready. Free memory (GB): 101.21/113.21, Free processors: 24/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
260409-02:20:15,272 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:20:15,276 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:20:15,278 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:20:15,278 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm".
260409-02:20:15,282 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/ants_transform-segmentation-to-qsm".
260409-02:20:15,283 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm".
260409-02:20:15,311 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_nilearn_t1w-seg-resampled" <qsmxt.interfaces.nipype_interface_resample_like.ResampleLikeInterface>
260409-02:20:15,434 nipype.workflow INFO:
[Node] Executing "ants_transform-segmentation-to-qsm" <nipype.interfaces.ants.resampling.ApplyTransforms>
260409-02:20:17,177 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 0 jobs ready. Free memory (GB): 97.21/113.21, Free processors: 22/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
260409-02:20:17,996 nipype.workflow INFO:
[Node] Finished "ants_transform-segmentation-to-qsm", elapsed time 2.560962s.
260409-02:20:19,178 nipype.workflow INFO:
[Job 40] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.ants_transform-segmentation-to-qsm).
260409-02:20:19,181 nipype.workflow INFO:
[MultiProc] Running 2 tasks, and 0 jobs ready. Free memory (GB): 99.21/113.21, Free processors: 23/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
260409-02:20:25,314 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_nilearn_t1w-seg-resampled", elapsed time 10.001752s.
260409-02:20:27,187 nipype.workflow INFO:
[Job 39] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.nibabel_numpy_nilearn_t1w-seg-resampled).
260409-02:20:27,190 nipype.workflow INFO:
[MultiProc] Running 1 tasks, and 1 jobs ready. Free memory (GB): 101.21/113.21, Free processors: 24/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
260409-02:20:27,274 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.copyfile".
260409-02:20:27,278 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.copyfile" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqmp2ragehighres0p5isoslab_T1w/copyfile".
260409-02:20:27,279 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.copyfile".
260409-02:20:27,281 nipype.workflow INFO:
[Node] Executing "copyfile" <qsmxt.interfaces.nipype_interface_copyfile.DynamicCopyFiles>
260409-02:20:27,679 nipype.workflow INFO:
[Node] Finished "copyfile", elapsed time 0.397366s.
260409-02:20:29,189 nipype.workflow INFO:
[Job 48] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqmp2ragehighres0p5isoslab_T1w.copyfile).
260409-02:20:29,191 nipype.workflow INFO:
[MultiProc] Running 1 tasks, and 0 jobs ready. Free memory (GB): 101.21/113.21, Free processors: 24/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1
260409-02:25:20,617 nipype.workflow INFO:
[Node] Finished "fastsurfer_segment-t1", elapsed time 505.55027s.
260409-02:25:21,619 nipype.workflow INFO:
[Job 22] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.fastsurfer_segment-t1).
260409-02:25:21,621 nipype.workflow INFO:
[MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 113.21/113.21, Free processors: 32/32.
260409-02:25:21,687 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.numpy_numpy_nibabel_mgz2nii".
260409-02:25:21,691 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.numpy_numpy_nibabel_mgz2nii" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/numpy_numpy_nibabel_mgz2nii".
260409-02:25:21,692 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.numpy_numpy_nibabel_mgz2nii".
260409-02:25:21,695 nipype.workflow INFO:
[Node] Executing "numpy_numpy_nibabel_mgz2nii" <qsmxt.interfaces.nipype_interface_mgz2nii.Mgz2NiiInterface>
260409-02:25:22,39 nipype.workflow INFO:
[Node] Finished "numpy_numpy_nibabel_mgz2nii", elapsed time 0.342405s.
260409-02:25:23,622 nipype.workflow INFO:
[Job 33] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.numpy_numpy_nibabel_mgz2nii).
260409-02:25:23,623 nipype.workflow INFO:
[MultiProc] Running 0 tasks, and 2 jobs ready. Free memory (GB): 113.21/113.21, Free processors: 32/32.
260409-02:25:23,697 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:25:23,700 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:25:23,701 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled".
260409-02:25:23,703 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_transform-segmentation-to-qsm".
260409-02:25:23,707 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_transform-segmentation-to-qsm" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/ants_transform-segmentation-to-qsm".
260409-02:25:23,708 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_transform-segmentation-to-qsm".
260409-02:25:23,721 nipype.workflow INFO:
[Node] Executing "nibabel_numpy_nilearn_t1w-seg-resampled" <qsmxt.interfaces.nipype_interface_resample_like.ResampleLikeInterface>
260409-02:25:23,739 nipype.workflow INFO:
[Node] Executing "ants_transform-segmentation-to-qsm" <nipype.interfaces.ants.resampling.ApplyTransforms>
260409-02:25:25,566 nipype.workflow INFO:
[Node] Finished "ants_transform-segmentation-to-qsm", elapsed time 1.825409s.
260409-02:25:25,626 nipype.workflow INFO:
[Job 43] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.ants_transform-segmentation-to-qsm).
260409-02:25:25,628 nipype.workflow INFO:
[MultiProc] Running 1 tasks, and 1 jobs ready. Free memory (GB): 111.21/113.21, Free processors: 31/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
260409-02:25:25,695 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists2".
260409-02:25:25,697 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists2" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/combine_lists2".
260409-02:25:25,698 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists2".
260409-02:25:25,700 nipype.workflow INFO:
[Node] Executing "combine_lists2" <nipype.interfaces.utility.wrappers.Function>
260409-02:25:25,702 nipype.workflow INFO:
[Node] Finished "combine_lists2", elapsed time 0.000652s.
260409-02:25:27,628 nipype.workflow INFO:
[Job 51] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.combine_lists2).
260409-02:25:27,630 nipype.workflow INFO:
[MultiProc] Running 1 tasks, and 1 jobs ready. Free memory (GB): 111.21/113.21, Free processors: 31/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
260409-02:25:27,707 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.create_permutations".
260409-02:25:27,710 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.create_permutations" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/create_permutations".
260409-02:25:27,713 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.create_permutations".
260409-02:25:27,716 nipype.workflow INFO:
[Node] Executing "create_permutations" <nipype.interfaces.utility.wrappers.Function>
260409-02:25:27,718 nipype.workflow INFO:
[Node] Finished "create_permutations", elapsed time 0.000966s.
260409-02:25:29,630 nipype.workflow INFO:
[Job 71] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.create_permutations).
260409-02:25:29,632 nipype.workflow INFO:
[MultiProc] Running 1 tasks, and 1 jobs ready. Free memory (GB): 111.21/113.21, Free processors: 31/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
260409-02:25:31,633 nipype.workflow INFO:
[MultiProc] Running 1 tasks, and 4 jobs ready. Free memory (GB): 111.21/113.21, Free processors: 31/32.
Currently running:
* qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled
260409-02:25:31,735 nipype.workflow INFO:
[Node] Outdated cache found for "_nibabel_numpy_analyse-qsm0".
260409-02:25:31,738 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm0" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm0".
260409-02:25:31,739 nipype.workflow INFO:
[Node] Outdated cache found for "_nibabel_numpy_analyse-qsm0".
260409-02:25:31,742 nipype.workflow INFO:
[Node] Executing "_nibabel_numpy_analyse-qsm0" <qsmxt.interfaces.nipype_interface_analyse.AnalyseInterface>
260409-02:25:31,742 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm3" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm3".
260409-02:25:31,745 nipype.workflow INFO:
[Node] Executing "_nibabel_numpy_analyse-qsm3" <qsmxt.interfaces.nipype_interface_analyse.AnalyseInterface>
260409-02:25:31,739 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm1".
260409-02:25:31,742 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm2" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm2".
260409-02:25:31,794 nipype.workflow INFO:
[Node] Executing "_nibabel_numpy_analyse-qsm1" <qsmxt.interfaces.nipype_interface_analyse.AnalyseInterface>
260409-02:25:31,795 nipype.workflow INFO:
[Node] Executing "_nibabel_numpy_analyse-qsm2" <qsmxt.interfaces.nipype_interface_analyse.AnalyseInterface>
260409-02:25:33,71 nipype.workflow INFO:
[Node] Finished "nibabel_numpy_nilearn_t1w-seg-resampled", elapsed time 9.348794s.
260409-02:25:33,635 nipype.workflow INFO:
[Job 42] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_nilearn_t1w-seg-resampled).
260409-02:25:33,637 nipype.workflow INFO:
[MultiProc] Running 4 tasks, and 0 jobs ready. Free memory (GB): 105.21/113.21, Free processors: 28/32.
Currently running:
* _nibabel_numpy_analyse-qsm3
* _nibabel_numpy_analyse-qsm2
* _nibabel_numpy_analyse-qsm1
* _nibabel_numpy_analyse-qsm0
260409-02:26:29,593 nipype.workflow INFO:
[Node] Finished "_nibabel_numpy_analyse-qsm2", elapsed time 57.796354s.
260409-02:26:29,734 nipype.workflow INFO:
[Job 82] Completed (_nibabel_numpy_analyse-qsm2).
260409-02:26:29,736 nipype.workflow INFO:
[MultiProc] Running 3 tasks, and 0 jobs ready. Free memory (GB): 107.21/113.21, Free processors: 29/32.
Currently running:
* _nibabel_numpy_analyse-qsm3
* _nibabel_numpy_analyse-qsm1
* _nibabel_numpy_analyse-qsm0
260409-02:26:34,677 nipype.workflow INFO:
[Node] Finished "_nibabel_numpy_analyse-qsm3", elapsed time 62.930457s.
260409-02:26:34,979 nipype.workflow INFO:
[Node] Finished "_nibabel_numpy_analyse-qsm0", elapsed time 63.235843s.
260409-02:26:35,399 nipype.workflow INFO:
[Node] Finished "_nibabel_numpy_analyse-qsm1", elapsed time 63.602155s.
260409-02:26:35,744 nipype.workflow INFO:
[Job 80] Completed (_nibabel_numpy_analyse-qsm0).
260409-02:26:35,745 nipype.workflow INFO:
[Job 81] Completed (_nibabel_numpy_analyse-qsm1).
260409-02:26:35,746 nipype.workflow INFO:
[Job 83] Completed (_nibabel_numpy_analyse-qsm3).
260409-02:26:35,747 nipype.workflow INFO:
[MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 113.21/113.21, Free processors: 32/32.
260409-02:26:35,827 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_analyse-qsm".
260409-02:26:35,831 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_analyse-qsm".
260409-02:26:35,836 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm0" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm0".
260409-02:26:35,841 nipype.workflow INFO:
[Node] Cached "_nibabel_numpy_analyse-qsm0" - collecting precomputed outputs
260409-02:26:35,842 nipype.workflow INFO:
[Node] "_nibabel_numpy_analyse-qsm0" found cached.
260409-02:26:35,844 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm1" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm1".
260409-02:26:35,851 nipype.workflow INFO:
[Node] Cached "_nibabel_numpy_analyse-qsm1" - collecting precomputed outputs
260409-02:26:35,852 nipype.workflow INFO:
[Node] "_nibabel_numpy_analyse-qsm1" found cached.
260409-02:26:35,854 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm2" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm2".
260409-02:26:35,856 nipype.workflow INFO:
[Node] Cached "_nibabel_numpy_analyse-qsm2" - collecting precomputed outputs
260409-02:26:35,856 nipype.workflow INFO:
[Node] "_nibabel_numpy_analyse-qsm2" found cached.
260409-02:26:35,865 nipype.workflow INFO:
[Node] Setting-up "_nibabel_numpy_analyse-qsm3" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/nibabel_numpy_analyse-qsm/mapflow/_nibabel_numpy_analyse-qsm3".
260409-02:26:35,867 nipype.workflow INFO:
[Node] Cached "_nibabel_numpy_analyse-qsm3" - collecting precomputed outputs
260409-02:26:35,868 nipype.workflow INFO:
[Node] "_nibabel_numpy_analyse-qsm3" found cached.
260409-02:26:37,746 nipype.workflow INFO:
[Job 73] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.nibabel_numpy_analyse-qsm).
260409-02:26:37,748 nipype.workflow INFO:
[MultiProc] Running 0 tasks, and 1 jobs ready. Free memory (GB): 113.21/113.21, Free processors: 32/32.
260409-02:26:37,852 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copyfile".
260409-02:26:37,855 nipype.workflow INFO:
[Node] Setting-up "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copyfile" in "/home/jovyan/workspace/books/examples/quantitative_imaging/bids/derivatives/workflow/qsmxt-workflow/sub-2/ses-20170705/qsm_acq-acqqsmp21mmisote20_T2starw/copyfile".
260409-02:26:37,856 nipype.workflow INFO:
[Node] Outdated cache found for "qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copyfile".
260409-02:26:37,859 nipype.workflow INFO:
[Node] Executing "copyfile" <qsmxt.interfaces.nipype_interface_copyfile.DynamicCopyFiles>
260409-02:26:38,70 nipype.workflow INFO:
[Node] Finished "copyfile", elapsed time 0.208956s.
260409-02:26:39,747 nipype.workflow INFO:
[Job 75] Completed (qsmxt-workflow.sub-2.ses-20170705.qsm_acq-acqqsmp21mmisote20_T2starw.copyfile).
260409-02:26:39,749 nipype.workflow INFO:
[MultiProc] Running 0 tasks, and 0 jobs ready. Free memory (GB): 113.21/113.21, Free processors: 32/32.
[INFO]: Warnings occurred!
[INFO]: Finished
View results#
Let’s have a look at the generated qsm folder:
!tree bids/derivatives/qsmxt-*-* --dirsfirst
bids/derivatives/qsmxt-2026-04-09-013659
├── sub-1
│ └── ses-20170705
│ ├── anat
│ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-orig_dseg.nii
│ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-qsm_dseg.nii
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.json
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.nii
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-orig_dseg.nii
│ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_dseg.nii
│ └── extra_data
│ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_desc-t1w-to-qsm_transform.mat
│ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-t1w-to-qsm_transform.mat
│ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_qsm-analysis.csv
├── sub-2
│ └── ses-20170705
│ ├── anat
│ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-orig_dseg.nii
│ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-qsm_dseg.nii
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.json
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.nii
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-orig_dseg.nii
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_dseg.nii
│ └── extra_data
│ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_desc-t1w-to-qsm_transform.mat
│ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-t1w-to-qsm_transform.mat
│ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_qsm-analysis.csv
├── command.txt
├── pypeline.log
├── qsmxt.log
├── references.txt
└── settings.json
bids/derivatives/qsmxt-2026-04-09-020519
├── sub-1
│ └── ses-20170705
│ ├── anat
│ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-orig_dseg.nii
│ │ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-qsm_dseg.nii
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.json
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.nii
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-orig_dseg.nii
│ │ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_dseg.nii
│ └── extra_data
│ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_qsm-analysis
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-qsmxt-2026-04-09-013659_Chimap_sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_desc-qsmxt-2026-04-09-013659_dseg_analysis.csv
│ │ ├── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-qsmxt-2026-04-09-013659_Chimap_sub-1_ses-20170705_space-qsm_desc-qsmxt-2026-04-09-020519_dseg_analysis.csv
│ │ ├── sub-1_ses-20170705_desc-qsmxt-2026-04-09-020519_Chimap_sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_desc-qsmxt-2026-04-09-013659_dseg_analysis.csv
│ │ └── sub-1_ses-20170705_desc-qsmxt-2026-04-09-020519_Chimap_sub-1_ses-20170705_space-qsm_desc-qsmxt-2026-04-09-020519_dseg_analysis.csv
│ ├── sub-1_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_desc-t1w-to-qsm_transform.mat
│ └── sub-1_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-t1w-to-qsm_transform.mat
├── sub-2
│ └── ses-20170705
│ ├── anat
│ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-orig_dseg.nii
│ │ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_space-qsm_dseg.nii
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.json
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_Chimap.nii
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-orig_dseg.nii
│ │ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_dseg.nii
│ └── extra_data
│ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_qsm-analysis
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-qsmxt-2026-04-09-013659_Chimap_sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_desc-qsmxt-2026-04-09-013659_dseg_analysis.csv
│ │ ├── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-qsmxt-2026-04-09-013659_Chimap_sub-2_ses-20170705_space-qsm_desc-qsmxt-2026-04-09-020519_dseg_analysis.csv
│ │ ├── sub-2_ses-20170705_desc-qsmxt-2026-04-09-020519_Chimap_sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_space-qsm_desc-qsmxt-2026-04-09-013659_dseg_analysis.csv
│ │ └── sub-2_ses-20170705_desc-qsmxt-2026-04-09-020519_Chimap_sub-2_ses-20170705_space-qsm_desc-qsmxt-2026-04-09-020519_dseg_analysis.csv
│ ├── sub-2_ses-20170705_acq-acqmp2ragehighres0p5isoslab_T1w_desc-t1w-to-qsm_transform.mat
│ └── sub-2_ses-20170705_acq-acqqsmp21mmisote20_T2starw_desc-t1w-to-qsm_transform.mat
├── command.txt
├── pypeline.log
├── qsmxt.log
├── references.txt
└── settings.json
20 directories, 52 files
The references.txt file contains a list of all the algorithms used and relevant citations:
!cat bids/derivatives/qsmxt-*-*/references.txt
== References ==
- QSMxT: Stewart AW, Robinson SD, O'Brien K, et al. QSMxT: Robust masking and artifact reduction for quantitative susceptibility mapping. Magnetic Resonance in Medicine. 2022;87(3):1289-1300. doi:10.1002/mrm.29048
- QSMxT: Stewart AW, Bollman S, et al. QSMxT/QSMxT. GitHub; 2022. https://github.com/QSMxT/QSMxT
- Python package - Nipype: Gorgolewski K, Burns C, Madison C, et al. Nipype: A Flexible, Lightweight and Extensible Neuroimaging Data Processing Framework in Python. Frontiers in Neuroinformatics. 2011;5. Accessed April 20, 2022. doi:10.3389/fninf.2011.00013
- Brain extraction: Smith SM. Fast robust automated brain extraction. Human Brain Mapping. 2002;17(3):143-155. doi:10.1002/hbm.10062
- Brain extraction: Liangfu Chen. liangfu/bet2 - Standalone Brain Extraction Tool. GitHub; 2015. https://github.com/liangfu/bet2
- Unwrapping algorithm - ROMEO: Dymerska B, Eckstein K, Bachrata B, et al. Phase unwrapping with a rapid opensource minimum spanning tree algorithm (ROMEO). Magnetic Resonance in Medicine. 2021;85(4):2294-2308. doi:10.1002/mrm.28563
- Background field removal - V-SHARP: Wu B, Li W, Guidon A et al. Whole brain susceptibility mapping using compressed sensing. Magnetic resonance in medicine. 2012 Jan;67(1):137-47. doi:10.1002/mrm.23000
- QSM algorithm - RTS: Kames C, Wiggermann V, Rauscher A. Rapid two-step dipole inversion for susceptibility mapping with sparsity priors. Neuroimage. 2018 Feb 15;167:276-83. doi:10.1016/j.neuroimage.2017.11.018
- Julia package - QSM.jl: kamesy. GitHub; 2022. https://github.com/kamesy/QSM.jl
- Julia package - MriResearchTools: Eckstein K. korbinian90/MriResearchTools.jl. GitHub; 2022. https://github.com/korbinian90/MriResearchTools.jl
- Python package - nibabel: Brett M, Markiewicz CJ, Hanke M, et al. nipy/nibabel. GitHub; 2019. https://github.com/nipy/nibabel
- Python package - scipy: Virtanen P, Gommers R, Oliphant TE, et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat Methods. 2020;17(3):261-272. doi:10.1038/s41592-019-0686-2
- Python package - numpy: Harris CR, Millman KJ, van der Walt SJ, et al. Array programming with NumPy. Nature. 2020;585(7825):357-362. doi:10.1038/s41586-020-2649-2
- FastSurfer: Henschel L, Conjeti S, Estrada S, Diers K, Fischl B, Reuter M. FastSurfer - A fast and accurate deep learning based neuroimaging pipeline. NeuroImage. 2020;219:117012. doi:10.1016/j.neuroimage.2020.117012
- ANTs: Avants BB, Tustison NJ, Johnson HJ. Advanced Normalization Tools. GitHub; 2022. https://github.com/ANTsX/ANTs
== References ==
- QSMxT: Stewart AW, Robinson SD, O'Brien K, et al. QSMxT: Robust masking and artifact reduction for quantitative susceptibility mapping. Magnetic Resonance in Medicine. 2022;87(3):1289-1300. doi:10.1002/mrm.29048
- QSMxT: Stewart AW, Bollman S, et al. QSMxT/QSMxT. GitHub; 2022. https://github.com/QSMxT/QSMxT
- Python package - Nipype: Gorgolewski K, Burns C, Madison C, et al. Nipype: A Flexible, Lightweight and Extensible Neuroimaging Data Processing Framework in Python. Frontiers in Neuroinformatics. 2011;5. Accessed April 20, 2022. doi:10.3389/fninf.2011.00013
- Brain extraction: Smith SM. Fast robust automated brain extraction. Human Brain Mapping. 2002;17(3):143-155. doi:10.1002/hbm.10062
- Brain extraction: Liangfu Chen. liangfu/bet2 - Standalone Brain Extraction Tool. GitHub; 2015. https://github.com/liangfu/bet2
- Unwrapping algorithm - ROMEO: Dymerska B, Eckstein K, Bachrata B, et al. Phase unwrapping with a rapid opensource minimum spanning tree algorithm (ROMEO). Magnetic Resonance in Medicine. 2021;85(4):2294-2308. doi:10.1002/mrm.28563
- Background field removal - V-SHARP: Wu B, Li W, Guidon A et al. Whole brain susceptibility mapping using compressed sensing. Magnetic resonance in medicine. 2012 Jan;67(1):137-47. doi:10.1002/mrm.23000
- QSM algorithm - RTS: Kames C, Wiggermann V, Rauscher A. Rapid two-step dipole inversion for susceptibility mapping with sparsity priors. Neuroimage. 2018 Feb 15;167:276-83. doi:10.1016/j.neuroimage.2017.11.018
- Julia package - QSM.jl: kamesy. GitHub; 2022. https://github.com/kamesy/QSM.jl
- Julia package - MriResearchTools: Eckstein K. korbinian90/MriResearchTools.jl. GitHub; 2022. https://github.com/korbinian90/MriResearchTools.jl
- Python package - nibabel: Brett M, Markiewicz CJ, Hanke M, et al. nipy/nibabel. GitHub; 2019. https://github.com/nipy/nibabel
- Python package - scipy: Virtanen P, Gommers R, Oliphant TE, et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat Methods. 2020;17(3):261-272. doi:10.1038/s41592-019-0686-2
- Python package - numpy: Harris CR, Millman KJ, van der Walt SJ, et al. Array programming with NumPy. Nature. 2020;585(7825):357-362. doi:10.1038/s41586-020-2649-2
- FastSurfer: Henschel L, Conjeti S, Estrada S, Diers K, Fischl B, Reuter M. FastSurfer - A fast and accurate deep learning based neuroimaging pipeline. NeuroImage. 2020;219:117012. doi:10.1016/j.neuroimage.2020.117012
- ANTs: Avants BB, Tustison NJ, Johnson HJ. Advanced Normalization Tools. GitHub; 2022. https://github.com/ANTsX/ANTs
QSM results#
Let’s view one of the QSM results:
show_nii(glob("bids/derivatives/qsmxt-*/sub-*/ses-*/anat/*Chimap.nii*")[0], title="Chimap", figsize=(10,8), cmap='gray', vmin=-0.1, vmax=0.1)
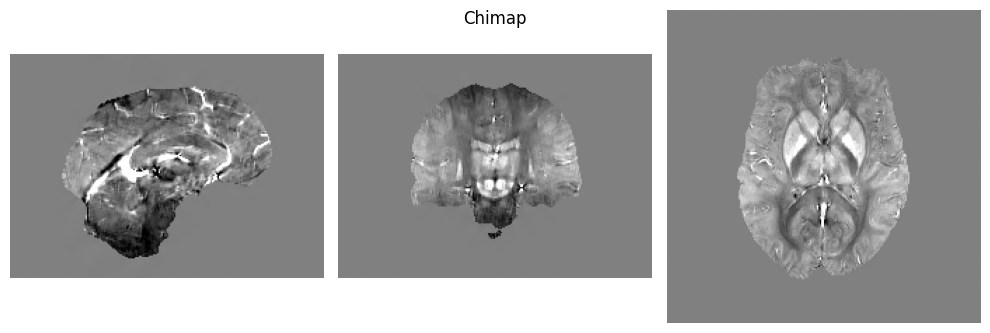
Segmentations#
Segmentations are generated in both the QSM space and the T1-weighted space. Transformations are also made available.
show_nii(glob("bids/derivatives/qsmxt-*/sub-*/ses-*/anat/*qsm_dseg.nii*")[0], title="Segmentation", figsize=(10,8), cmap='Set3', vmin=0, vmax=96)
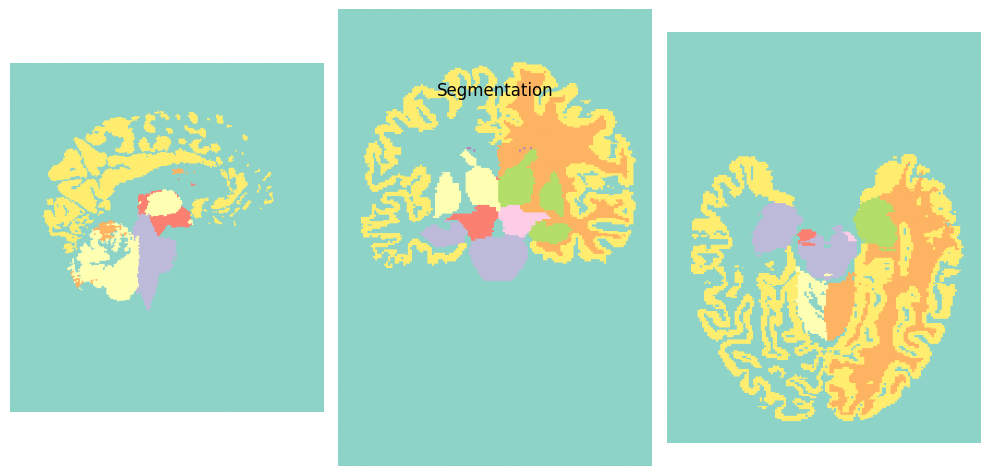
Analysis CSVs#
CSV files have been exported containing susceptibility values in regions of interest for each subject.
Here we will load the CSVs, inspect the data and generate figures:
# import modules
import pandas as pd
import seaborn as sns
The raw CSV files use names from FreeSurfer as exported by FastSurfer. The full list of regions is available here.
pd.read_csv(glob("bids/derivatives/qsmxt-*-*/sub*/ses*/extra*/*.csv")[0])
| roi | num_voxels | min | max | median | mean | std | |
|---|---|---|---|---|---|---|---|
| 0 | 3rd-Ventricle | 587 | -0.234321 | 0.054945 | -0.018367 | -0.021842 | 0.033371 |
| 1 | 4th-Ventricle | 1543 | -0.126117 | 0.052944 | -0.044647 | -0.045172 | 0.030772 |
| 2 | Brain-Stem | 18516 | -0.750579 | 0.102235 | -0.046106 | -0.042326 | 0.028948 |
| 3 | CSF | 780 | -1.393693 | 0.945752 | 0.030650 | 0.033611 | 0.142992 |
| 4 | Left-Accumbens-area | 518 | -0.141920 | 0.128334 | 0.028726 | 0.023371 | 0.031773 |
| ... | ... | ... | ... | ... | ... | ... | ... |
| 91 | ctx-rh-superiorfrontal | 29675 | -0.103077 | 0.249447 | -0.003901 | -0.009051 | 0.023885 |
| 92 | ctx-rh-superiorparietal | 12154 | -0.084225 | 0.271532 | -0.015429 | -0.014625 | 0.021848 |
| 93 | ctx-rh-superiortemporal | 16362 | -0.090982 | 0.248737 | 0.006287 | 0.012667 | 0.024258 |
| 94 | ctx-rh-supramarginal | 12103 | -0.056940 | 0.239688 | 0.018211 | 0.019522 | 0.022483 |
| 95 | ctx-rh-transversetemporal | 1237 | -0.004704 | 0.171156 | 0.034416 | 0.035389 | 0.015418 |
96 rows × 7 columns
We will select a subset of these ROIs and give them more readable names:
# define regions of interest
# see https://github.com/QSMxT/QSMxT/blob/main/qsmxt/aseg_labels.csv for a full list
rois = {
"Thalamus" : [9, 10, 48, 49],
"Pallidum" : [12, 13, 52, 53],
"Caudate" : [11, 50],
"Putamen" : [12, 51],
"Brain stem" : [16],
"CSF" : [24, 122, 257, 701],
"White matter" : [2, 7, 41, 46, 177]
}
roi_names = { value: key for key in rois for value in rois[key] }
roi_ids = [value for roi in rois.values() for value in roi]
# load a reconstruction
qsm_img = nib.load(glob("bids/derivatives/qsmxt-*-*/sub*/ses*/anat*/*Chimap.nii*")[0])
seg_img = nib.load(glob("bids/derivatives/qsmxt-*-*/sub*/ses*/anat*/*space-qsm*dseg.nii*")[0])
from nilearn.image import resample_to_img
# Resample the segmentation image to match the QSM image space
# Use 'nearest' interpolation to preserve the integer labels of the segmentation.
print("Resampling segmentation to match QSM dimensions...")
seg_resampled_img = resample_to_img(seg_img, qsm_img, interpolation='nearest')
print("Resampling complete.")
Resampling segmentation to match QSM dimensions...
Resampling complete.
# Get data arrays
qsm = qsm_img.get_fdata()
seg = seg_resampled_img.get_fdata()
# Check shapes to confirm they match
print(f"Shape of QSM data: {qsm.shape}")
print(f"Shape of resampled segmentation data: {seg.shape}")
if qsm.shape != seg.shape:
raise ValueError("FATAL: Shapes not match after resampling. Check your input files.")
Shape of QSM data: (224, 224, 160)
Shape of resampled segmentation data: (224, 224, 160)
# --- Flatten the arrays now that they have the same dimensions ---
qsm = qsm.flatten()
seg = seg.flatten()
# retain only the rois
qsm = qsm[np.isin(seg, roi_ids)]
seg = seg[np.isin(seg, roi_ids)]
# convert to a dataframe for plotting purposes
seg = pd.Series(seg).map(roi_names)
data = pd.DataFrame({ 'qsm' : qsm, 'seg' : seg })
# summarise data by region including the average and standard deviation
data.groupby('seg')['qsm'].agg(['mean', 'std']).sort_values('mean').round(decimals=3)
| mean | std | |
|---|---|---|
| seg | ||
| CSF | -0.040 | 0.030 |
| Thalamus | -0.010 | 0.022 |
| Brain stem | -0.004 | 0.017 |
| Putamen | -0.003 | 0.012 |
| White matter | -0.003 | 0.027 |
| Caudate | -0.002 | 0.013 |
| Pallidum | -0.001 | 0.008 |
medians = data.groupby('seg')['qsm'].median().sort_values()
order = medians.index
# plot
fig, ax = plt.subplots(figsize=(5,4))
sns.boxplot(data=data, y='qsm', x='seg', fliersize=0, color='lightblue', order=order, ax=ax)
ax.set_xticks(range(len(ax.get_xticks()))) # Set tick positions explicitly
ax.set_xticklabels(ax.get_xticklabels(), ha='right', rotation=45)
ax.set_ylim(-0.2, 0.3)
ax.axhline(y=0, color='pink', linestyle='-', linewidth=1, zorder=-1)
ax.set_xlabel("Region of interest")
ax.set_ylabel("Susceptibility (ppm)")
ax.set_title("QSM")
plt.show()
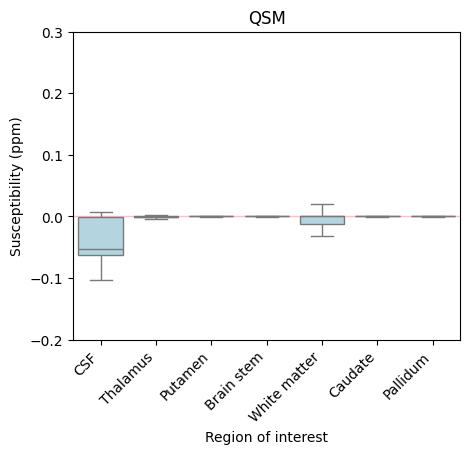
Dependencies in Jupyter/Python#
Using the package watermark to document system environment and software versions used in this notebook, alongside the Neurodesktop version extracted from the
JUPYTER_IMAGEorNEURODESKTOP_VERSIONenvironment variables.
import os
%load_ext watermark
%watermark
%watermark --iversions
neurodesktop_version = (
os.environ.get('JUPYTER_IMAGE', '').split(':')[-1] or
os.environ.get('NEURODESKTOP_VERSION', 'unknown')
)
print(f"Neurodesktop version: {neurodesktop_version}")
Last updated: 2026-04-09T02:26:47.093318+00:00
Python implementation: CPython
Python version : 3.13.9
IPython version : 9.7.0
Compiler : GCC 14.3.0
OS : Linux
Release : 5.15.0-171-generic
Machine : x86_64
Processor : x86_64
CPU cores : 32
Architecture: 64bit
ipyniivue : 2.4.4
json : 2.0.9
matplotlib: 3.10.8
nibabel : 5.3.3
nilearn : 0.13.1
numpy : 2.3.5
pandas : 2.3.3
seaborn : 0.13.2
Neurodesktop version: 2025-12-20
