Neurodesk Tools Demo#
Shell We Begin?#
Welcome to this Neurodesk demonstration notebook - a guide to the rich ecosystem of tools, workflows, and visualization strategies available on the Neurodesk platform. From command-line tools and pipelines to modular Python interfaces, this notebook provides an overview of setting up pipelines, running individual processing steps, and inspecting outputs, from shell and cell.
Use this notebook to:
Explore a selection of popular neuroimaging tools (e.g., FSL, AFNI, FreeSurfer, MRIQC)
Learn how to run CLI-based commands, set up workflows, and execute Pythonic processing steps interactively
Visualize outputs and quality control reports directly in the notebook
Navigate to detailed, tool-specific notebooks for deeper exploration
Author: Monika Doerig
Date: 13 June 2025
License:
Note: If this notebook uses neuroimaging tools from Neurocontainers, those tools retain their original licenses. Please see Neurodesk citation guidelines for details.
Citation: For clarity and conciseness, references and citations supporting each tool or workflow are included in the linked notebooks themselves.
1. Load software tools and import Python libraries#
%%capture
!pip install fury==0.11.0 dipy==1.11.0 vtk==9.5.2 pybids==0.19.0 antspyx nilearn
import os
import nibabel as nib
from scipy import ndimage
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import transforms
import ants
from bids.layout import BIDSLayout, BIDSValidator
from ipyniivue import NiiVue
from IPython.display import Image, display
from IPython.core.display import SVG
from nipype.interfaces import fsl
from nipype.interfaces import afni
from dipy.core.gradients import gradient_table
from dipy.io import read_bvals_bvecs
from dipy.io.image import load_nifti
import dipy.reconst.dti as dti
from dipy.segment.mask import median_otsu
from dipy.data import get_sphere
from dipy.viz import actor, window
from nilearn.image.image import mean_img
from nilearn.plotting import plot_epi, plot_anat, plot_img, plot_roi, view_img
# We can use module to load the different software tools in a specific version
import module
await module.purge()
await module.load('afni/21.2.00')
await module.load('mriqc/24.0.2')
await module.load('ants/2.6.0')
await module.load('bidscoin/4.3.3')
await module.load('freesurfer/8.1.0')
await module.load('mrtrix3/3.0.4')
await module.load('fsl/6.0.7.16')
await module.list()
['afni/21.2.00',
'mriqc/24.0.2',
'ants/2.6.0',
'bidscoin/4.3.3',
'freesurfer/8.1.0',
'mrtrix3/3.0.4',
'fsl/6.0.7.16']
2. Data#
Access Open Data on OSF Using DataLad#
Note :This workflow requires the dataset to be available as a DataLad dataset on OSF (i.e. with DataLad metadata and annexed file storage).
# Download one subject of the Brain Tumor Connectomics Data in BIDS Format
PATTERN = "sub-CON02"
! datalad install https://github.com/OpenNeuroDatasets/ds001226.git
! cd ds001226 && datalad get $PATTERN
3. Bits about BIDS#
BIDS conversion with BIDScoin#
For more information on BIDS conversion using different software tools, dcm2niix, HeuDiConv, and BIDScoin, see this BIDS conversion example.
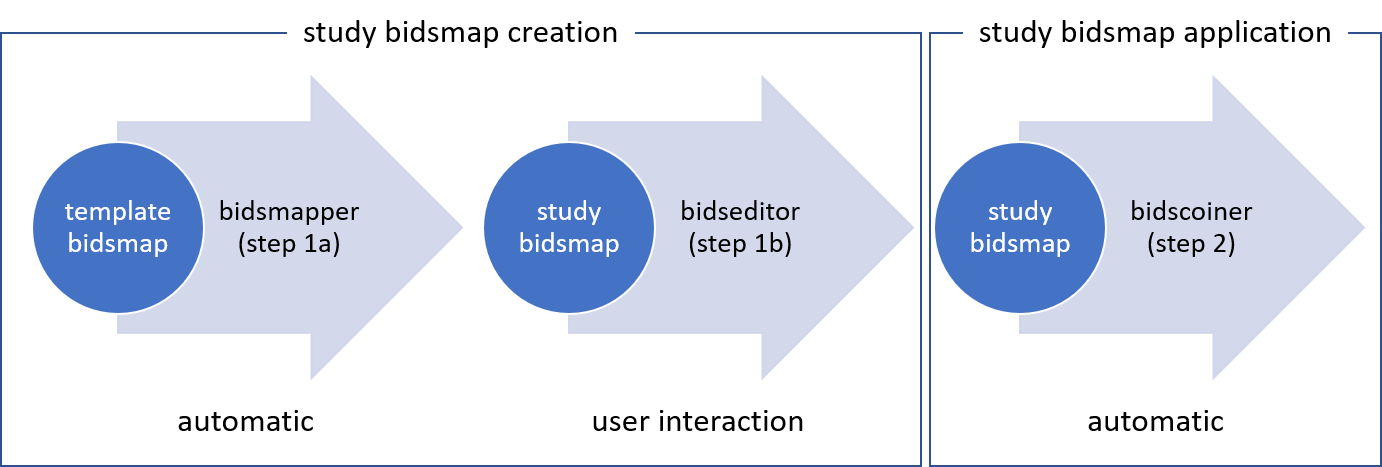
BIDScoin workflow:
BIDScoin requires that the source data repository follows a subject/[session]/data structure. The data folder can be organized in various DICOM layouts: DICOM series, DICOMDIR, or flat DICOMs.
To perform a BIDS conversion using BIDScoin:
Run
bidsmapperwith the--automatedflag to generate thebidsmapnon-interactively (this skips the manual editing step withbidseditor).Run
bidscoinerto convert the data using the generatedbidsmap.
! bidscoin --download . # Download the tutorial data (use a "." for the current folder or a pathname of choice to save it elsewhere)
INFO | Downloading the tutorial dataset...
bidscointutorial.tar.gz: 0.00B [00:00, ?B/s]
bidscointutorial.tar.gz: 0%| | 8.00k/192M [00:03<24:48:39, 2.25kB/s]
bidscointutorial.tar.gz: 0%| | 16.0k/192M [00:03<11:19:06, 4.94kB/s]
bidscointutorial.tar.gz: 0%| | 24.0k/192M [00:04<6:33:08, 8.53kB/s]
bidscointutorial.tar.gz: 0%| | 40.0k/192M [00:04<3:07:27, 17.9kB/s]
bidscointutorial.tar.gz: 0%| | 88.0k/192M [00:04<1:00:56, 55.0kB/s]
bidscointutorial.tar.gz: 0%| | 96.0k/192M [00:04<1:00:47, 55.1kB/s]
bidscointutorial.tar.gz: 0%| | 176k/192M [00:04<23:24, 143kB/s]
bidscointutorial.tar.gz: 0%| | 224k/192M [00:04<18:29, 181kB/s]
bidscointutorial.tar.gz: 0%| | 344k/192M [00:04<10:00, 335kB/s]
bidscointutorial.tar.gz: 0%| | 440k/192M [00:05<08:01, 417kB/s]
bidscointutorial.tar.gz: 0%| | 712k/192M [00:05<04:02, 826kB/s]
bidscointutorial.tar.gz: 0%| | 904k/192M [00:05<03:26, 968kB/s]
bidscointutorial.tar.gz: 1%| | 1.41M/192M [00:05<01:49, 1.83MB/s]
bidscointutorial.tar.gz: 1%|▏ | 1.77M/192M [00:05<01:37, 2.05MB/s]
bidscointutorial.tar.gz: 1%|▏ | 2.84M/192M [00:05<00:51, 3.85MB/s]
bidscointutorial.tar.gz: 2%|▎ | 3.55M/192M [00:05<00:46, 4.22MB/s]
bidscointutorial.tar.gz: 3%|▍ | 5.30M/192M [00:05<00:27, 6.99MB/s]
bidscointutorial.tar.gz: 3%|▍ | 6.42M/192M [00:06<00:26, 7.32MB/s]
bidscointutorial.tar.gz: 4%|▌ | 7.50M/192M [00:06<00:25, 7.69MB/s]
bidscointutorial.tar.gz: 5%|▋ | 9.20M/192M [00:06<00:21, 9.11MB/s]
bidscointutorial.tar.gz: 5%|▋ | 10.2M/192M [00:06<00:21, 8.80MB/s]
bidscointutorial.tar.gz: 6%|▊ | 11.8M/192M [00:06<00:19, 9.77MB/s]
bidscointutorial.tar.gz: 7%|▉ | 12.9M/192M [00:06<00:20, 9.34MB/s]
bidscointutorial.tar.gz: 7%|█ | 14.3M/192M [00:06<00:19, 9.74MB/s]
bidscointutorial.tar.gz: 8%|█▏ | 15.6M/192M [00:07<00:18, 9.77MB/s]
bidscointutorial.tar.gz: 9%|█▏ | 16.9M/192M [00:07<00:18, 9.73MB/s]
bidscointutorial.tar.gz: 9%|█▎ | 18.0M/192M [00:07<00:19, 9.49MB/s]
bidscointutorial.tar.gz: 10%|█▍ | 19.5M/192M [00:07<00:18, 9.82MB/s]
bidscointutorial.tar.gz: 11%|█▍ | 20.4M/192M [00:07<00:19, 9.10MB/s]
bidscointutorial.tar.gz: 12%|█▌ | 22.1M/192M [00:07<00:17, 10.1MB/s]
bidscointutorial.tar.gz: 12%|█▋ | 22.9M/192M [00:07<00:19, 9.05MB/s]
bidscointutorial.tar.gz: 13%|█▊ | 24.7M/192M [00:08<00:16, 10.3MB/s]
bidscointutorial.tar.gz: 13%|█▊ | 25.5M/192M [00:08<00:19, 9.06MB/s]
bidscointutorial.tar.gz: 14%|█▉ | 27.2M/192M [00:08<00:16, 10.2MB/s]
bidscointutorial.tar.gz: 15%|██ | 28.1M/192M [00:08<00:18, 9.20MB/s]
bidscointutorial.tar.gz: 16%|██▏ | 29.8M/192M [00:08<00:16, 10.1MB/s]
bidscointutorial.tar.gz: 16%|██▏ | 30.6M/192M [00:08<00:18, 9.16MB/s]
bidscointutorial.tar.gz: 17%|██▎ | 32.3M/192M [00:08<00:16, 10.0MB/s]
bidscointutorial.tar.gz: 17%|██▍ | 33.1M/192M [00:09<00:18, 9.18MB/s]
bidscointutorial.tar.gz: 18%|██▌ | 34.8M/192M [00:09<00:16, 10.1MB/s]
bidscointutorial.tar.gz: 19%|██▌ | 35.6M/192M [00:09<00:17, 9.11MB/s]
bidscointutorial.tar.gz: 19%|██▋ | 37.3M/192M [00:09<00:16, 10.1MB/s]
bidscointutorial.tar.gz: 20%|██▊ | 38.2M/192M [00:09<00:17, 9.16MB/s]
bidscointutorial.tar.gz: 21%|██▉ | 39.9M/192M [00:09<00:15, 10.0MB/s]
bidscointutorial.tar.gz: 21%|██▉ | 40.7M/192M [00:09<00:17, 9.21MB/s]
bidscointutorial.tar.gz: 22%|███ | 42.4M/192M [00:09<00:15, 9.97MB/s]
bidscointutorial.tar.gz: 23%|███▏ | 43.2M/192M [00:10<00:17, 9.10MB/s]
bidscointutorial.tar.gz: 23%|███▎ | 44.9M/192M [00:10<00:15, 10.1MB/s]
bidscointutorial.tar.gz: 24%|███▎ | 45.7M/192M [00:10<00:16, 9.09MB/s]
bidscointutorial.tar.gz: 25%|███▍ | 47.5M/192M [00:10<00:15, 10.1MB/s]
bidscointutorial.tar.gz: 25%|███▌ | 48.3M/192M [00:10<00:16, 9.09MB/s]
bidscointutorial.tar.gz: 26%|███▋ | 50.0M/192M [00:10<00:14, 10.1MB/s]
bidscointutorial.tar.gz: 26%|███▋ | 50.8M/192M [00:10<00:16, 9.11MB/s]
bidscointutorial.tar.gz: 27%|███▊ | 52.5M/192M [00:11<00:14, 10.1MB/s]
bidscointutorial.tar.gz: 28%|███▉ | 53.4M/192M [00:11<00:15, 9.16MB/s]
bidscointutorial.tar.gz: 29%|████ | 55.0M/192M [00:11<00:14, 10.0MB/s]
bidscointutorial.tar.gz: 29%|████ | 55.9M/192M [00:11<00:15, 9.15MB/s]
bidscointutorial.tar.gz: 30%|████▏ | 57.5M/192M [00:11<00:14, 9.96MB/s]
bidscointutorial.tar.gz: 30%|████▎ | 58.4M/192M [00:11<00:15, 9.21MB/s]
bidscointutorial.tar.gz: 31%|████▍ | 60.1M/192M [00:11<00:13, 10.0MB/s]
bidscointutorial.tar.gz: 32%|████▍ | 60.9M/192M [00:12<00:14, 9.16MB/s]
bidscointutorial.tar.gz: 33%|████▌ | 62.6M/192M [00:12<00:13, 10.0MB/s]
bidscointutorial.tar.gz: 33%|████▋ | 63.5M/192M [00:12<00:14, 9.19MB/s]
bidscointutorial.tar.gz: 34%|████▊ | 65.2M/192M [00:12<00:13, 10.0MB/s]
bidscointutorial.tar.gz: 34%|████▊ | 66.0M/192M [00:12<00:14, 9.13MB/s]
bidscointutorial.tar.gz: 35%|████▉ | 67.7M/192M [00:12<00:12, 10.1MB/s]
bidscointutorial.tar.gz: 36%|█████ | 68.5M/192M [00:12<00:14, 9.17MB/s]
bidscointutorial.tar.gz: 37%|█████▏ | 70.5M/192M [00:13<00:11, 10.6MB/s]
bidscointutorial.tar.gz: 37%|█████▏ | 71.1M/192M [00:13<00:14, 8.96MB/s]
bidscointutorial.tar.gz: 38%|█████▎ | 73.2M/192M [00:13<00:11, 11.0MB/s]
bidscointutorial.tar.gz: 38%|█████▍ | 73.7M/192M [00:13<00:13, 9.07MB/s]
bidscointutorial.tar.gz: 40%|█████▌ | 75.9M/192M [00:13<00:10, 11.1MB/s]
bidscointutorial.tar.gz: 40%|█████▌ | 76.4M/192M [00:13<00:13, 9.17MB/s]
bidscointutorial.tar.gz: 41%|█████▋ | 78.5M/192M [00:13<00:10, 10.9MB/s]
bidscointutorial.tar.gz: 41%|█████▊ | 79.1M/192M [00:13<00:12, 9.26MB/s]
bidscointutorial.tar.gz: 42%|█████▉ | 81.1M/192M [00:14<00:10, 10.7MB/s]
bidscointutorial.tar.gz: 43%|█████▉ | 81.8M/192M [00:14<00:12, 9.28MB/s]
bidscointutorial.tar.gz: 44%|██████ | 83.6M/192M [00:14<00:10, 10.5MB/s]
bidscointutorial.tar.gz: 44%|██████▏ | 84.4M/192M [00:14<00:12, 9.29MB/s]
bidscointutorial.tar.gz: 45%|██████▎ | 86.1M/192M [00:14<00:10, 10.3MB/s]
bidscointutorial.tar.gz: 45%|██████▎ | 87.0M/192M [00:14<00:11, 9.33MB/s]
bidscointutorial.tar.gz: 46%|██████▍ | 88.7M/192M [00:14<00:10, 10.4MB/s]
bidscointutorial.tar.gz: 47%|██████▌ | 89.5M/192M [00:15<00:11, 9.13MB/s]
bidscointutorial.tar.gz: 48%|██████▋ | 91.3M/192M [00:15<00:10, 10.4MB/s]
bidscointutorial.tar.gz: 48%|██████▋ | 92.0M/192M [00:15<00:11, 9.11MB/s]
bidscointutorial.tar.gz: 49%|██████▊ | 93.9M/192M [00:15<00:09, 10.4MB/s]
bidscointutorial.tar.gz: 49%|██████▉ | 94.6M/192M [00:15<00:11, 9.12MB/s]
bidscointutorial.tar.gz: 50%|███████ | 96.4M/192M [00:15<00:09, 10.3MB/s]
bidscointutorial.tar.gz: 51%|███████ | 97.2M/192M [00:15<00:10, 9.18MB/s]
bidscointutorial.tar.gz: 52%|███████▏ | 98.9M/192M [00:16<00:09, 10.2MB/s]
bidscointutorial.tar.gz: 52%|███████▎ | 99.8M/192M [00:16<00:10, 9.20MB/s]
bidscointutorial.tar.gz: 53%|███████▉ | 102M/192M [00:16<00:09, 10.3MB/s]
bidscointutorial.tar.gz: 53%|███████▉ | 102M/192M [00:16<00:10, 9.03MB/s]
bidscointutorial.tar.gz: 54%|████████▏ | 104M/192M [00:16<00:09, 10.0MB/s]
bidscointutorial.tar.gz: 55%|████████▏ | 105M/192M [00:16<00:09, 9.25MB/s]
bidscointutorial.tar.gz: 55%|████████▎ | 106M/192M [00:16<00:08, 9.97MB/s]
bidscointutorial.tar.gz: 56%|████████▍ | 107M/192M [00:17<00:09, 9.13MB/s]
bidscointutorial.tar.gz: 57%|████████▌ | 109M/192M [00:17<00:08, 9.96MB/s]
bidscointutorial.tar.gz: 57%|████████▌ | 110M/192M [00:17<00:09, 9.11MB/s]
bidscointutorial.tar.gz: 58%|████████▋ | 112M/192M [00:17<00:08, 10.1MB/s]
bidscointutorial.tar.gz: 59%|████████▊ | 112M/192M [00:17<00:09, 9.06MB/s]
bidscointutorial.tar.gz: 59%|████████▉ | 114M/192M [00:17<00:08, 10.1MB/s]
bidscointutorial.tar.gz: 60%|████████▉ | 115M/192M [00:17<00:08, 9.13MB/s]
bidscointutorial.tar.gz: 61%|█████████ | 117M/192M [00:17<00:07, 10.1MB/s]
bidscointutorial.tar.gz: 61%|█████████▏ | 117M/192M [00:18<00:08, 9.17MB/s]
bidscointutorial.tar.gz: 62%|█████████▎ | 119M/192M [00:18<00:07, 10.1MB/s]
bidscointutorial.tar.gz: 63%|█████████▍ | 120M/192M [00:18<00:08, 9.20MB/s]
bidscointutorial.tar.gz: 63%|█████████▌ | 122M/192M [00:18<00:07, 10.0MB/s]
bidscointutorial.tar.gz: 64%|█████████▌ | 123M/192M [00:18<00:07, 9.21MB/s]
bidscointutorial.tar.gz: 65%|█████████▋ | 124M/192M [00:18<00:07, 10.1MB/s]
bidscointutorial.tar.gz: 65%|█████████▊ | 125M/192M [00:18<00:07, 9.54MB/s]
bidscointutorial.tar.gz: 66%|█████████▉ | 127M/192M [00:19<00:06, 9.93MB/s]
bidscointutorial.tar.gz: 67%|█████████▉ | 128M/192M [00:19<00:07, 9.26MB/s]
bidscointutorial.tar.gz: 67%|██████████ | 129M/192M [00:19<00:06, 9.87MB/s]
bidscointutorial.tar.gz: 68%|██████████▏ | 130M/192M [00:19<00:07, 9.14MB/s]
bidscointutorial.tar.gz: 69%|██████████▎ | 132M/192M [00:19<00:06, 10.1MB/s]
bidscointutorial.tar.gz: 69%|██████████▎ | 133M/192M [00:19<00:06, 9.04MB/s]
bidscointutorial.tar.gz: 70%|██████████▌ | 134M/192M [00:19<00:05, 10.1MB/s]
bidscointutorial.tar.gz: 70%|██████████▌ | 135M/192M [00:20<00:06, 9.08MB/s]
bidscointutorial.tar.gz: 71%|██████████▋ | 137M/192M [00:20<00:05, 10.1MB/s]
bidscointutorial.tar.gz: 72%|██████████▊ | 138M/192M [00:20<00:06, 9.16MB/s]
bidscointutorial.tar.gz: 73%|██████████▉ | 139M/192M [00:20<00:05, 10.1MB/s]
bidscointutorial.tar.gz: 73%|██████████▉ | 140M/192M [00:20<00:05, 9.14MB/s]
bidscointutorial.tar.gz: 74%|███████████ | 142M/192M [00:20<00:05, 9.99MB/s]
bidscointutorial.tar.gz: 74%|███████████▏ | 143M/192M [00:20<00:05, 9.06MB/s]
bidscointutorial.tar.gz: 75%|███████████▎ | 144M/192M [00:21<00:04, 10.1MB/s]
bidscointutorial.tar.gz: 76%|███████████▎ | 145M/192M [00:21<00:05, 9.00MB/s]
bidscointutorial.tar.gz: 77%|███████████▍ | 147M/192M [00:21<00:04, 10.1MB/s]
bidscointutorial.tar.gz: 77%|███████████▌ | 148M/192M [00:21<00:05, 9.07MB/s]
bidscointutorial.tar.gz: 78%|███████████▋ | 150M/192M [00:21<00:04, 10.2MB/s]
bidscointutorial.tar.gz: 78%|███████████▊ | 150M/192M [00:21<00:04, 9.08MB/s]
bidscointutorial.tar.gz: 79%|███████████▉ | 152M/192M [00:21<00:04, 10.2MB/s]
bidscointutorial.tar.gz: 80%|███████████▉ | 153M/192M [00:21<00:04, 9.09MB/s]
bidscointutorial.tar.gz: 81%|████████████ | 155M/192M [00:22<00:03, 10.1MB/s]
bidscointutorial.tar.gz: 81%|████████████▏ | 155M/192M [00:22<00:04, 9.11MB/s]
bidscointutorial.tar.gz: 82%|████████████▎ | 157M/192M [00:22<00:03, 10.0MB/s]
bidscointutorial.tar.gz: 82%|████████████▎ | 158M/192M [00:22<00:03, 9.15MB/s]
bidscointutorial.tar.gz: 83%|████████████▍ | 160M/192M [00:22<00:03, 10.1MB/s]
bidscointutorial.tar.gz: 84%|████████████▌ | 160M/192M [00:22<00:03, 9.02MB/s]
bidscointutorial.tar.gz: 85%|████████████▋ | 162M/192M [00:22<00:03, 10.1MB/s]
bidscointutorial.tar.gz: 85%|████████████▋ | 163M/192M [00:23<00:03, 9.12MB/s]
bidscointutorial.tar.gz: 86%|████████████▊ | 165M/192M [00:23<00:02, 10.2MB/s]
bidscointutorial.tar.gz: 86%|████████████▉ | 166M/192M [00:23<00:03, 9.07MB/s]
bidscointutorial.tar.gz: 87%|█████████████ | 167M/192M [00:23<00:02, 10.2MB/s]
bidscointutorial.tar.gz: 88%|█████████████▏ | 168M/192M [00:23<00:02, 9.11MB/s]
bidscointutorial.tar.gz: 88%|█████████████▎ | 170M/192M [00:23<00:02, 10.1MB/s]
bidscointutorial.tar.gz: 89%|█████████████▎ | 171M/192M [00:23<00:02, 9.10MB/s]
bidscointutorial.tar.gz: 90%|█████████████▍ | 172M/192M [00:24<00:02, 10.0MB/s]
bidscointutorial.tar.gz: 90%|█████████████▌ | 173M/192M [00:24<00:02, 9.07MB/s]
bidscointutorial.tar.gz: 91%|█████████████▋ | 175M/192M [00:24<00:01, 10.1MB/s]
bidscointutorial.tar.gz: 92%|█████████████▋ | 176M/192M [00:24<00:01, 9.07MB/s]
bidscointutorial.tar.gz: 92%|█████████████▊ | 177M/192M [00:24<00:01, 10.1MB/s]
bidscointutorial.tar.gz: 93%|█████████████▉ | 178M/192M [00:24<00:01, 9.07MB/s]
bidscointutorial.tar.gz: 94%|██████████████ | 180M/192M [00:24<00:01, 10.1MB/s]
bidscointutorial.tar.gz: 94%|██████████████ | 181M/192M [00:25<00:01, 9.08MB/s]
bidscointutorial.tar.gz: 95%|██████████████▎| 182M/192M [00:25<00:00, 10.2MB/s]
bidscointutorial.tar.gz: 95%|██████████████▎| 183M/192M [00:25<00:01, 9.06MB/s]
bidscointutorial.tar.gz: 96%|██████████████▍| 185M/192M [00:25<00:00, 10.1MB/s]
bidscointutorial.tar.gz: 97%|██████████████▌| 186M/192M [00:25<00:00, 9.06MB/s]
bidscointutorial.tar.gz: 98%|██████████████▋| 187M/192M [00:25<00:00, 10.1MB/s]
bidscointutorial.tar.gz: 98%|██████████████▋| 188M/192M [00:25<00:00, 9.10MB/s]
bidscointutorial.tar.gz: 99%|██████████████▊| 190M/192M [00:25<00:00, 10.0MB/s]
bidscointutorial.tar.gz: 99%|██████████████▉| 191M/192M [00:26<00:00, 9.07MB/s]
bidscointutorial.tar.gz: 192MB [00:26, 7.69MB/s]
INFO | Unpacking the downloaded data in: /home/jovyan/workspace/books/examples/workflows
SUCCESS | Done
! bidsmapper -h
usage: bidsmapper [-h] [-b NAME] [-t NAME] [-p NAME [NAME ...]] [-n PREFIX]
[-m PREFIX] [-u PATTERN] [-s] [-a] [-f] [--no-update]
sourcefolder bidsfolder
The bidsmapper scans your source data repository to identify different data types by matching
them against the run-items in the template bidsmap. Once a match is found, a mapping to BIDS
output data types is made and the run-item is added to the study bidsmap. You can check and
edit these generated bids-mappings to your needs with the (automatically launched) bidseditor.
Re-run the bidsmapper whenever something was changed in your data acquisition protocol and
edit the new data type to your needs (your existing bidsmap will be re-used).
The bidsmapper uses plugins, as stored in the bidsmap['Options'], to do the actual work
positional arguments:
sourcefolder The study root folder containing the raw source data
folders
bidsfolder The destination folder with the (future) bids data and
the bidsfolder/code/bidscoin/bidsmap.yaml output file
options:
-h, --help show this help message and exit
-b NAME, --bidsmap NAME
The study bidsmap file with the mapping heuristics. If
the bidsmap filename is just the basename (i.e. no '/'
in the name) then it is assumed to be located in the
current directory or in bidsfolder/code/bidscoin.
Default: bidsmap.yaml
-t NAME, --template NAME
The bidsmap template file with the default heuristics
(this could be provided by your institute). If the
bidsmap filename is just the basename (i.e. no '/' in
the name) then it is assumed to be located in the
bidscoin config folder. Default: bidsmap_dccn
-p NAME [NAME ...], --plugins NAME [NAME ...]
List of plugins to be used. Default: the plugin list
of the study/template bidsmap)
-n PREFIX, --subprefix PREFIX
The prefix common for all the source subject-folders
(e.g. 'Pt' is the subprefix if subject folders are
named 'Pt018', 'Pt019', ...). Use '*' when your
subject folders do not have a prefix. Default: the
value of the study/template bidsmap, e.g. 'sub-'
-m PREFIX, --sesprefix PREFIX
The prefix common for all the source session-folders
(e.g. 'M_' is the subprefix if session folders are
named 'M_pre', 'M_post', ..). Use '*' when your
session folders do not have a prefix. Default: the
value of the study/template bidsmap, e.g. 'ses-'
-u PATTERN, --unzip PATTERN
Wildcard pattern to unpack tarball/zip-files in the
sub/ses sourcefolder that need to be unzipped (in a
tempdir) to make the data readable. Default: the value
of the study/template bidsmap
-s, --store Store provenance data samples in the
bidsfolder/code/provenance folder (useful for
inspecting e.g. zipped or transferred datasets)
-a, --automated Save the automatically generated bidsmap to disk and
without interactively tweaking it with the bidseditor
-f, --force Discard the previously saved bidsmap and logfile
--no-update Do not update any sub/sesprefixes in or prepend the
sourcefolder name to the <<filepath:regex>> expression
that extracts the subject/session labels. This is
normally done to make the extraction more robust, but
could cause problems for certain use cases
examples:
bidsmapper myproject/raw myproject/bids
bidsmapper myproject/raw myproject/bids -t bidsmap_custom # Uses a template bidsmap of choice
bidsmapper myproject/raw myproject/bids -p nibabel2bids # Uses a plugin of choice
bidsmapper myproject/raw myproject/bids -u '*.tar.gz' # Unzip tarball sourcefiles
! bidsmapper ./bidscointutorial/raw ./bidscointutorial/bids --automated
! bidscoiner ./bidscointutorial/raw ./bidscointutorial/bids
INFO |
INFO | -------------- START BIDSmapper ------------
INFO | >>> bidsmapper sourcefolder=/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw bidsfolder=/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids bidsmap=bidsmap.yaml template=/home/jovyan/.bidscoin/4.3.3+qt5/templates/bidsmap_dccn.yaml plugins=[] subprefix=None sesprefix=None store=False force=False
INFO | Reading: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/code/bidscoin/bidsmap.yaml
INFO | Checking the bidsmap run-items:
SUCCESS | All run-items in the bidsmap are valid
INFO | Reading: /home/jovyan/.bidscoin/4.3.3+qt5/templates/bidsmap_dccn.yaml
INFO | Checking the bidsmap run-items:
SUCCESS | All datatypes and options in the template bidsmap are valid
INFO | Mapping: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-001/ses-01 (subject 1/2)
VERBOSE | Executing plugin: dcm2niix2bids -> /home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-001/ses-01
0%| | 0/2 [00:00<?, ?subject/s]
INFO | Mapping: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01 (subject 2/2)
INFO | Detected a flat data-structure in: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01
INFO | Making temporary copy: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01 -> /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01
50%|████████████████████ | 1/2 [00:00<00:00, 2.88subject/s]
INFO | >> Sorting: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01 (712 files)
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/060-cmrr_2p5iso_mb3me3_TR1500
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/002-AAHead_Scout_32ch-head
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/007-t1_mprage_sag_ipat2_1p0iso
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/048-cmrr_2p4iso_mb8_TR0700
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/050-field_map_2p4iso
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/049-field_map_2p4iso
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/062-field_map_2p5iso
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/061-field_map_2p5iso
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/059-cmrr_2p5iso_mb3me3_TR1500_SBRef
50%|████████████████████ | 1/2 [00:00<00:00, 2.88subject/s]
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/047-cmrr_2p4iso_mb8_TR0700_SBRef
50%|████████████████████ | 1/2 [00:00<00:00, 2.88subject/s]
VERBOSE | Creating: /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/001-localizer_32ch-head
50%|████████████████████ | 1/2 [00:00<00:00, 2.88subject/s]
VERBOSE | Executing plugin: dcm2niix2bids -> /tmp/tmp39v29has/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01
50%|████████████████████ | 1/2 [00:01<00:00, 2.88subject/s]
INFO | Checking the bidsmap run-items:
SUCCESS | All run-items in the bidsmap are valid
INFO | bids-validator 1.14.6 test results (* = in .bidsignore):
INFO | False: exclude/sub-DICOM_task-localizer32chhead_acq-localizer32chhead_echo-1_GR.*
INFO | False: exclude/sub-DICOM_task-AAHeadScout32chhead_acq-AAHeadScout32chhead_echo-1_GR.*
SUCCESS | All generated bidsnames are BIDS-valid
INFO | Writing bidsmap to: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/code/bidscoin/bidsmap.yaml
INFO | -------------- FINISHED! -------------------
INFO |
SUCCESS | No BIDScoin errors or warnings were reported
INFO |
INFO | For the complete log see: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/code/bidscoin/bidsmapper.log
NB: That folder may contain privacy sensitive information, e.g. pathnames in logfiles and provenance data samples
INFO |
INFO | -------------- START BIDScoiner 4.3.3+qt5: BIDS 1.9.0 ------------
INFO | >>> bidscoiner sourcefolder=/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw bidsfolder=/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids participant=None force=False bidsmap=bidsmap.yaml
INFO | Reading: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/code/bidscoin/bidsmap.yaml
INFO | Checking the bidsmap run-items:
SUCCESS | All run-items in the bidsmap are valid
INFO | ------------------- Subject 1/2 -------------------
INFO | >>> Skipping processed session: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/sub-001/ses-01 already has {'func', 'anat', 'fmap'} data (you can carefully use the -f option to overrule)
INFO | ------------------- Subject 2/2 -------------------
0%| | 0/2 [00:00<?, ?subject/s]
INFO | Detected a flat data-structure in: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01
INFO | Making temporary copy: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01 -> /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01
0%| | 0/2 [00:00<?, ?subject/s]
INFO | >> Sorting: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01 (712 files)
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/060-cmrr_2p5iso_mb3me3_TR1500
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/002-AAHead_Scout_32ch-head
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/007-t1_mprage_sag_ipat2_1p0iso
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/048-cmrr_2p4iso_mb8_TR0700
0%| | 0/2 [00:00<?, ?subject/s]
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/050-field_map_2p4iso
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/049-field_map_2p4iso
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/062-field_map_2p5iso
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/061-field_map_2p5iso
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/059-cmrr_2p5iso_mb3me3_TR1500_SBRef
0%| | 0/2 [00:00<?, ?subject/s]
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/047-cmrr_2p4iso_mb8_TR0700_SBRef
0%| | 0/2 [00:00<?, ?subject/s]
VERBOSE | Creating: /tmp/tmp3b8ym5bw/home/jovyan/workspace/books/examples/workflows/bidscointutorial/raw/sub-002/ses-01/001-localizer_32ch-head
0%| | 0/2 [00:00<?, ?subject/s]
INFO | >>> Skipping processed session: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/sub-002/ses-01 already has {'func', 'anat', 'fmap'} data (you can carefully use the -f option to overrule)
INFO | -------------- FINISHED! ------------
INFO |
SUCCESS | No BIDScoin errors or warnings were reported
INFO |
INFO | For the complete log see: /home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/code/bidscoin/bidscoiner.log
NB: That folder may contain privacy sensitive information, e.g. pathnames in logfiles and provenance data samples
!tree -L 4 ./bidscointutorial/bids
./bidscointutorial/bids
├── README
├── code
│ └── bidscoin
│ ├── bidscoiner.errors
│ ├── bidscoiner.log
│ ├── bidscoiner.tsv
│ ├── bidsmap.yaml
│ ├── bidsmapper.errors
│ └── bidsmapper.log
├── dataset_description.json
├── participants.json
├── participants.tsv
├── sub-001
│ └── ses-01
│ ├── anat
│ │ ├── sub-001_ses-01_acq-t1mpragesagipat21p0iso_T1w.json
│ │ └── sub-001_ses-01_acq-t1mpragesagipat21p0iso_T1w.nii.gz
│ ├── fmap
│ │ ├── sub-001_ses-01_acq-fieldmap2p4iso_magnitude1.json
│ │ ├── sub-001_ses-01_acq-fieldmap2p4iso_magnitude1.nii.gz
│ │ ├── sub-001_ses-01_acq-fieldmap2p4iso_magnitude2.json
│ │ ├── sub-001_ses-01_acq-fieldmap2p4iso_magnitude2.nii.gz
│ │ ├── sub-001_ses-01_acq-fieldmap2p4iso_phasediff.json
│ │ ├── sub-001_ses-01_acq-fieldmap2p4iso_phasediff.nii.gz
│ │ ├── sub-001_ses-01_acq-fieldmap2p5iso_magnitude1.json
│ │ ├── sub-001_ses-01_acq-fieldmap2p5iso_magnitude1.nii.gz
│ │ ├── sub-001_ses-01_acq-fieldmap2p5iso_magnitude2.json
│ │ ├── sub-001_ses-01_acq-fieldmap2p5iso_magnitude2.nii.gz
│ │ ├── sub-001_ses-01_acq-fieldmap2p5iso_phasediff.json
│ │ └── sub-001_ses-01_acq-fieldmap2p5iso_phasediff.nii.gz
│ ├── func
│ │ ├── sub-001_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_bold.json
│ │ ├── sub-001_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_bold.nii.gz
│ │ ├── sub-001_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_sbref.json
│ │ ├── sub-001_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_sbref.nii.gz
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_bold.json
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_bold.nii.gz
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_sbref.json
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_sbref.nii.gz
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_bold.json
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_bold.nii.gz
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_sbref.json
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_sbref.nii.gz
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_bold.json
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_bold.nii.gz
│ │ ├── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_sbref.json
│ │ └── sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_sbref.nii.gz
│ └── sub-001_ses-01_scans.tsv
└── sub-002
└── ses-01
├── anat
│ ├── sub-002_ses-01_acq-t1mpragesagipat21p0iso_T1w.json
│ └── sub-002_ses-01_acq-t1mpragesagipat21p0iso_T1w.nii.gz
├── fmap
│ ├── sub-002_ses-01_acq-fieldmap2p4iso_magnitude1.json
│ ├── sub-002_ses-01_acq-fieldmap2p4iso_magnitude1.nii.gz
│ ├── sub-002_ses-01_acq-fieldmap2p4iso_magnitude2.json
│ ├── sub-002_ses-01_acq-fieldmap2p4iso_magnitude2.nii.gz
│ ├── sub-002_ses-01_acq-fieldmap2p4iso_phasediff.json
│ ├── sub-002_ses-01_acq-fieldmap2p4iso_phasediff.nii.gz
│ ├── sub-002_ses-01_acq-fieldmap2p5iso_magnitude1.json
│ ├── sub-002_ses-01_acq-fieldmap2p5iso_magnitude1.nii.gz
│ ├── sub-002_ses-01_acq-fieldmap2p5iso_magnitude2.json
│ ├── sub-002_ses-01_acq-fieldmap2p5iso_magnitude2.nii.gz
│ ├── sub-002_ses-01_acq-fieldmap2p5iso_phasediff.json
│ └── sub-002_ses-01_acq-fieldmap2p5iso_phasediff.nii.gz
├── func
│ ├── sub-002_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_bold.json
│ ├── sub-002_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_bold.nii.gz
│ ├── sub-002_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_sbref.json
│ ├── sub-002_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_sbref.nii.gz
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_bold.json
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_bold.nii.gz
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_sbref.json
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_sbref.nii.gz
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_bold.json
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_bold.nii.gz
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_sbref.json
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_sbref.nii.gz
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_bold.json
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_bold.nii.gz
│ ├── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_sbref.json
│ └── sub-002_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_sbref.nii.gz
└── sub-002_ses-01_scans.tsv
13 directories, 72 files
pyBIDS#
PyBIDS is a Python package that makes it easier to query, summarize, and manage BIDS-formatted datasets. It integrates smoothly with neuroimaging tools like Nipype and Nilearn, which support the BIDS standard natively. Here, you’ll finda more comprehensive example.
BIDSLayout#
layout = BIDSLayout("./bidscointutorial/bids")
all_files = layout.get()
print("There are {} files in the layout.".format(len(all_files)))
print("\nThe first 3 files are:")
all_files[:5]
There are 66 files in the layout.
The first 3 files are:
[<BIDSJSONFile filename='/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/dataset_description.json'>,
<BIDSJSONFile filename='/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/participants.json'>,
<BIDSDataFile filename='/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/participants.tsv'>,
<BIDSFile filename='/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/README'>,
<BIDSJSONFile filename='/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/sub-001/ses-01/anat/sub-001_ses-01_acq-t1mpragesagipat21p0iso_T1w.json'>]
# Retrieve filenames of all BOLD runs for subject 01
bold_run = layout.get(subject='001', suffix='bold', extension='nii.gz', return_type='filename')
bold_run
['/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/sub-001/ses-01/func/sub-001_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_bold.nii.gz',
'/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/sub-001/ses-01/func/sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-1_bold.nii.gz',
'/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/sub-001/ses-01/func/sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-2_bold.nii.gz',
'/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/sub-001/ses-01/func/sub-001_ses-01_task-cmrr2p5isomb3me3TR1500_dir-AP_echo-3_bold.nii.gz']
layout.get_task()
['cmrr2p4isomb8TR0700', 'cmrr2p5isomb3me3TR1500']
The BIDSFile#
# Pick the 11th file in the dataset
bf = layout.get()[19]
# Print it
bf
<BIDSImageFile filename='/home/jovyan/workspace/books/examples/workflows/bidscointutorial/bids/sub-001/ses-01/func/sub-001_ses-01_task-cmrr2p4isomb8TR0700_dir-AP_bold.nii.gz'>
bf.get_entities()
{'datatype': 'func',
'direction': 'AP',
'extension': '.nii.gz',
'session': '01',
'subject': '001',
'suffix': 'bold',
'task': 'cmrr2p4isomb8TR0700'}
# Print all metadata of this file
bf.get_metadata()
{'AcquisitionMatrixPE': 88,
'AcquisitionNumber': 1,
'AcquisitionTime': '10:52:57.307500',
'BandwidthPerPixelPhaseEncode': 17.756,
'BaseResolution': 88,
'BidsGuess': ['func', '_acq-epfid2m8_dir-AP_run-48_bold'],
'BodyPartExamined': 'BRAIN',
'CoilCombinationMethod': 'Sum of Squares',
'ConsistencyInfo': 'N4_VE11C_LATEST_20160120',
'ConversionSoftware': 'dcm2niix',
'ConversionSoftwareVersion': 'v1.0.20240202',
'DerivedVendorReportedEchoSpacing': 0.000639989,
'DeviceSerialNumber': '66068',
'DwellTime': 2.8e-06,
'EchoTime': 0.039,
'EchoTrainLength': 88,
'EffectiveEchoSpacing': 0.000639989,
'FlipAngle': 52,
'ImageComments': 'Unaliased MB8/PE3/LB SENSE1',
'ImageOrientationPatientDICOM': [1, 0, 0, 0, 1, 0],
'ImageOrientationText': 'Tra',
'ImageType': ['ORIGINAL', 'PRIMARY', 'M', 'MB', 'ND', 'NORM', 'MOSAIC'],
'ImagingFrequency': 123.253552,
'InPlanePhaseEncodingDirectionDICOM': 'COL',
'InstitutionAddress': 'Kapittelweg 29,Nijmegen,Gelderland,NL,6525EN',
'InstitutionName': 'Donders Centre for Cognitive Neuroimaging',
'InstitutionalDepartmentName': 'Department',
'MRAcquisitionType': '2D',
'MagneticFieldStrength': 3,
'Manufacturer': 'Siemens',
'ManufacturersModelName': 'Prisma',
'MatrixCoilMode': 'SENSE',
'Modality': 'MR',
'MultibandAccelerationFactor': 8,
'NonlinearGradientCorrection': False,
'PartialFourier': 1,
'PatientPosition': 'HFS',
'PercentPhaseFOV': 100,
'PercentSampling': 100,
'PhaseEncodingDirection': 'j-',
'PhaseEncodingSteps': 88,
'PhaseResolution': 1,
'PixelBandwidth': 2030,
'ProcedureStepDescription': 'PauGaa^DCCN',
'ProtocolName': 'cmrr_2p4iso_mb8_TR0700',
'PulseSequenceDetails': '%CustomerSeq%\\cmrr_mbep2d_bold',
'ReceiveCoilActiveElements': 'HEA;HEP',
'ReceiveCoilName': 'Head_32',
'ReconMatrixPE': 88,
'RepetitionTime': 0.7,
'SAR': 0.0382286,
'ScanOptions': 'FS',
'ScanningSequence': 'EP',
'SequenceName': 'epfid2d1_88',
'SequenceVariant': 'SK\\SS',
'SeriesDescription': 'cmrr_2p4iso_mb8_TR0700',
'SeriesNumber': 48,
'ShimSetting': [3614, 2402, -2974, -439, 73, 372, 51, -15],
'SliceThickness': 2.4,
'SliceTiming': [0,
0.255,
0.51,
0.085,
0.34,
0.595,
0.17,
0.425,
0,
0.255,
0.51,
0.085,
0.34,
0.595,
0.17,
0.425,
0,
0.255,
0.51,
0.085,
0.34,
0.595,
0.17,
0.425,
0,
0.255,
0.51,
0.085,
0.34,
0.595,
0.17,
0.425,
0,
0.255,
0.51,
0.085,
0.34,
0.595,
0.17,
0.425,
0,
0.255,
0.51,
0.085,
0.34,
0.595,
0.17,
0.425,
0,
0.255,
0.51,
0.085,
0.34,
0.595,
0.17,
0.425,
0,
0.255,
0.51,
0.085,
0.34,
0.595,
0.17,
0.425],
'SoftwareVersions': 'syngo MR E11',
'SpacingBetweenSlices': 2.4,
'StationName': 'PRISMA',
'TaskName': 'cmrr_2p4iso_mb8_TR0700',
'TotalReadoutTime': 0.055679,
'TxRefAmp': 241.323,
'WipMemBlock': '46420e6b-85a4-4239-95df-b37f61133eb4||Sequence: R016 ve11c/master r/e634e98; Dec 19 2017 11:00:25 by eja'}
The BIDSValidator#
# When using the bids validator, the filepath MUST be relative to the top level bids directory
validator = BIDSValidator()
validator.is_bids('/sub-002/ses-01/anat/sub-002_ses-01_acq-t1mpragesagipat21p0iso_T1w.nii.gz')
True
4. Toolbox Tour#
4.1. Workflows#
Several neuroimaging tools provide streamlined, reproducible pipelines that build on widely used software packages such as FSL, AFNI, ANTs, and SPM. Some notable examples, like fMRIPrep and MRIQC, are high-level, preconfigured pipelines built on Nipype, a flexible Python workflow engine designed for creating custom neuroimaging processing chains.
fMRIPrep: A Preprocessing Pipeline for fMRI Data#
fMRIPrep offers a robust BIDS-compatible pipeline for preprocessing fMRI data with minimal manual intervention, as demonstrated in the fMRIPrep example notebook.
MRIQC: Quality control#
MRIQC provides automated quality assessment metrics for structural and functional MRI data, generating comprehensive reports to identify potential issues before analysis. For a complete walkthrough and more detailed example, see the full MRIQC example.
!mriqc --help
usage: mriqc [-h] [--version] [-v] [--species {human,rat}]
[--participant-label PARTICIPANT_LABEL [PARTICIPANT_LABEL ...]]
[--bids-filter-file PATH] [--session-id [SESSION_ID ...]]
[--run-id [RUN_ID ...]] [--task-id [TASK_ID ...]]
[-m [{T1w,T2w,bold,dwi} ...]] [--dsname DSNAME]
[--bids-database-dir PATH] [--bids-database-wipe]
[--no-datalad-get] [--nprocs NPROCS]
[--omp-nthreads OMP_NTHREADS] [--mem MEMORY_GB] [--testing] [-f]
[--pdb] [-w WORK_DIR] [--verbose-reports] [--reports-only]
[--write-graph] [--dry-run] [--resource-monitor]
[--use-plugin USE_PLUGIN] [--crashfile-format {txt,pklz}]
[--no-sub] [--email EMAIL] [--webapi-url WEBAPI_URL]
[--webapi-port WEBAPI_PORT] [--upload-strict] [--notrack]
[--ants-float] [--ants-settings ANTS_SETTINGS]
[--min-dwi-length MIN_LEN_DWI] [--min-bold-length MIN_LEN_BOLD]
[--fft-spikes-detector] [--fd_thres FD_THRES] [--deoblique]
[--despike] [--start-idx START_IDX] [--stop-idx STOP_IDX]
bids_dir output_dir {participant,group} [{participant,group} ...]
MRIQC 24.1.0.dev0+gd5b13cb5.d20240826 Automated Quality Control and visual
reports for Quality Assessment of structural (T1w, T2w) and functional MRI of
the brain. IMPORTANT: Anonymized quality metrics (IQMs) will be submitted to
MRIQC's metrics repository. Submission of IQMs can be disabled using the
``--no-sub`` argument. Please visit
https://mriqc.readthedocs.io/en/latest/dsa.html to revise MRIQC's Data Sharing
Agreement.
positional arguments:
bids_dir The root folder of a BIDS valid dataset (sub-XXXXX
folders should be found at the top level in this
folder).
output_dir The directory where the output files should be stored.
If you are running group level analysis this folder
should be prepopulated with the results of the
participant level analysis.
{participant,group} Level of the analysis that will be performed. Multiple
participant level analyses can be run independently
(in parallel) using the same output_dir.
options:
-h, --help show this help message and exit
--version show program's version number and exit
-v, --verbose Increases log verbosity for each occurrence, debug
level is -vvv. (default: 0)
--species {human,rat}
Use appropriate template for population (default:
human)
Options for filtering BIDS queries:
--participant-label PARTICIPANT_LABEL [PARTICIPANT_LABEL ...], --participant_label PARTICIPANT_LABEL [PARTICIPANT_LABEL ...], --participant-labels PARTICIPANT_LABEL [PARTICIPANT_LABEL ...], --participant_labels PARTICIPANT_LABEL [PARTICIPANT_LABEL ...]
A space delimited list of participant identifiers or a
single identifier (the sub- prefix can be removed).
(default: None)
--bids-filter-file PATH
a JSON file describing custom BIDS input filter using
pybids {<suffix>:{<entity>:<filter>,...},...}
(https://github.com/bids-standard/pybids/blob/master/b
ids/layout/config/bids.json) (default: None)
--session-id [SESSION_ID ...]
Filter input dataset by session ID. (default: None)
--run-id [RUN_ID ...]
DEPRECATED - This argument will be disabled. Use
``--bids-filter-file`` instead. (default: None)
--task-id [TASK_ID ...]
Filter input dataset by task ID. (default: None)
-m [{T1w,T2w,bold,dwi} ...], --modalities [{T1w,T2w,bold,dwi} ...]
Filter input dataset by MRI type. (default: ('T1w',
'T2w', 'bold', 'dwi'))
--dsname DSNAME A dataset name. (default: None)
--bids-database-dir PATH
Path to an existing PyBIDS database folder, for faster
indexing (especially useful for large datasets).
(default: None)
--bids-database-wipe Wipe out previously existing BIDS indexing caches,
forcing re-indexing. (default: False)
--no-datalad-get Disable attempting to get remote files in DataLad
datasets. (default: True)
Options to handle performance:
--nprocs NPROCS, --n_procs NPROCS, --n_cpus NPROCS, -n-cpus NPROCS
Maximum number of simultaneously running parallel
processes executed by *MRIQC* (e.g., several instances
of ANTs' registration). However, when ``--nprocs`` is
greater or equal to the ``--omp-nthreads`` option, it
also sets the maximum number of threads that
simultaneously running processes may aggregate
(meaning, with ``--nprocs 16 --omp-nthreads 8`` a
maximum of two 8-CPU-threaded processes will be
running at a given time). Under this mode of
operation, ``--nprocs`` sets the maximum number of
processors that can be assigned work within an *MRIQC*
job, which includes all the processors used by
currently running single- and multi-threaded
processes. If ``None``, the number of CPUs available
will be automatically assigned (which may not be what
you want in, e.g., shared systems like a HPC cluster.
(default: None)
--omp-nthreads OMP_NTHREADS, --ants-nthreads OMP_NTHREADS
Maximum number of threads that multi-threaded
processes executed by *MRIQC* (e.g., ANTs'
registration) can use. If ``None``, the number of CPUs
available will be automatically assigned (which may
not be what you want in, e.g., shared systems like a
HPC cluster. (default: None)
--mem MEMORY_GB, --mem_gb MEMORY_GB, --mem-gb MEMORY_GB
Upper bound memory limit for MRIQC processes.
(default: None)
--testing Use testing settings for a minimal footprint.
(default: False)
-f, --float32 Cast the input data to float32 if it's represented in
higher precision (saves space and improves
performance). (default: True)
--pdb Open Python debugger (pdb) on exceptions. (default:
False)
Instrumental options:
-w WORK_DIR, --work-dir WORK_DIR
Path where intermediate results should be stored.
(default:
/home/jovyan/workspace/books/examples/workflows/work)
--verbose-reports
--reports-only
--write-graph Write workflow graph. (default: False)
--dry-run Do not run the workflow. (default: False)
--resource-monitor, --profile
Hook up the resource profiler callback to nipype.
(default: False)
--use-plugin USE_PLUGIN
Nipype plugin configuration file. (default: None)
--crashfile-format {txt,pklz}
Nipype crashfile format (default: txt)
--no-sub Turn off submission of anonymized quality metrics to
MRIQC's metrics repository. (default: False)
--email EMAIL Email address to include with quality metric
submission. (default: )
--webapi-url WEBAPI_URL
IP address where the MRIQC WebAPI is listening.
(default: None)
--webapi-port WEBAPI_PORT
Port where the MRIQC WebAPI is listening. (default:
None)
--upload-strict Upload will fail if upload is strict. (default: False)
--notrack Opt-out of sending tracking information of this run to
the NiPreps developers. This information helps to
improve MRIQC and provides an indicator of real world
usage crucial for obtaining funding. (default: False)
Specific settings for ANTs:
--ants-float Use float number precision on ANTs computations.
(default: False)
--ants-settings ANTS_SETTINGS
Path to JSON file with settings for ANTs. (default:
None)
Diffusion MRI workflow configuration:
--min-dwi-length MIN_LEN_DWI
Drop DWI runs with fewer orientations than this
threshold. (default: 7)
Functional MRI workflow configuration:
--min-bold-length MIN_LEN_BOLD
Drop BOLD runs with fewer time points than this
threshold. (default: 5)
--fft-spikes-detector
Turn on FFT based spike detector (slow). (default:
False)
--fd_thres FD_THRES Threshold on framewise displacement estimates to
detect outliers. (default: 0.2)
--deoblique Deoblique the functional scans during head motion
correction preprocessing. (default: False)
--despike Despike the functional scans during head motion
correction preprocessing. (default: False)
--start-idx START_IDX
DEPRECATED Initial volume in functional timeseries
that should be considered for preprocessing. (default:
None)
--stop-idx STOP_IDX DEPRECATED Final volume in functional timeseries that
should be considered for preprocessing. (default:
None)
%%bash
# Set the number of threads for ITK (Image Processing Toolkit) to use
export ITK_GLOBAL_DEFAULT_NUMBER_OF_THREADS=6
# Set the directory for matplotlib configuration to avoid conflicts
export MPLCONFIGDIR=~/matplotlib-mpldir
# Run MRIQC
mriqc ds001226 \
MRIQC \
participant \
--participant-label sub-CON02 \
--no-sub \
--work-dir MRIQC_workdir \
--nprocs 10 \
--mem-gb 10 \
--verbose-reports \
-v
------------------------------------------------------------------
Running MRIQC version 24.1.0.de
v0+gd5b13cb5.d20240826
----------------------------------------------------------------
NOTICE
Copyright © The NiPreps Developers.
This product includes software developed by
the NiPrep
s Community (https://nipreps.org/).
Portions of this software were developed at the Department
of
Psychology at Stanford University, Stanford, CA, US.
This software contains code ultimatel
y derived from the
PCP Quality Assessment Protocol (QAP;
http://preprocessed-connectomes-project
.org/quality-assessment-protocol)
by C. Craddock, S. Giavasis, D. Clark, Z. Shezhad, and J. Pellma
n.
This software is also distributed as a Docker container image.
The bootstrapping file for
the image ("Dockerfile") is licensed
under the MIT License.
-----------------------------------
-----------------------------
* BIDS dataset path: /home/jovyan/workspace/books/examples/workflow
s/ds001226.
* Output folder: MRIQC.
* Analysis levels: ['participant'].
------------------------
------------------------------------------
2026-04-10 23:31:05 | IMPORTANT | mriqc | DataLad dataset
identified, attempting to `datalad get` unavailable files.
action summary:
get (notneeded: 4)
2026-04-10 23:31:05 | IMPORTANT | mriqc
| Extracting metadata and entities for 1 input runs of modality 't1w'...
2026-04-10 23:31:05 | IMPORTANT | mriqc | File size ('t1w
'): 0.01|0.01 GB [maximum|average].
2026-04-10 23:31:05 | IMPORTANT | mriqc | Extracting meta
data and entities for 1 input runs of modality 'bold'...
2026-04-10 23:31:06 | IMPORTANT | mriqc | File size ('bol
d'): 0.04|0.04 GB [maximum|average].
2026-04-10 23:31:06 | IMPORTANT | mriqc
| Extracting metadata and entities for 2 input runs of modality 'dwi'...
2026-04-10 23:31:06 | WARNING | nipype.workflow | 1 cannot be proc
essed (too short or too long)
2026-04-10 23:31:06 | IMPORTANT | mriqc | File size ('dwi
'): 0.04|0.04 GB [maximum|average].
2026-04-10 23:31:06 | INFO | mriqc | MRIQC config fil
e: /home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/config-20260410-233102_0780926c-793
c-4ef5-b154-1c12d857b918.toml.
2026-04-10 23:31:15 | IMPORTANT | mriqc | Building MRIQC'
s workflows...
2026-04-10 23:31:16 | INFO | nipype.workflow | Building functio
nal MRIQC workflow for 1 BOLD runs..
2026-04-10 23:31:17 | INFO | nipype.workflow | Building diffusi
on MRIQC workflow for 1 NIfTI files..
2026-04-10 23:31:17 | INFO | nipype.workflow | Building anatomi
cal MRIQC workflow for 1 NIfTI files..
2026-04-10 23:31:25 | IMPORTANT | mriqc | Workflow buildi
ng finished (exit code 0).
2026-04-10 23:31:32 | INFO | nipype.workflow | Workflow mriqc_w
f settings: ['check', 'execution', 'logging', 'monitoring']
2026-04-10 23:31:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.load_bmat".
2026-04-10 23:31:33 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.dwiMRIQC.load_bmat" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mr
iqc_wf/dwiMRIQC/edbb42c7bf5828aa552d68da572c7aebc01fdb50/load_bmat".
2026-04-10 23:31:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.load_bmat".
2026-04-10 23:31:33 | INFO | nipype.workflow | [Node] Executing
"load_bmat" <mriqc.interfaces.diffusion.ReadDWIMetadata>
2026-04-10 23:31:33 | INFO | nipype.workflow | [Node] Finished
"load_bmat", elapsed time 0.363733s.
2026-04-10 23:31:37 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_background".
2026-04-10 23:31:37 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_background" in "/home/jovyan/workspace/books/examples
/workflows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/
ds_report_background".
2026-04-10 23:31:37 | INFO | nipype.workflow 0
m| [Node] Outdated cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_background".
2026-04-10 23:31:37 | INFO | nipype.workflow | [Node] Executing
"ds_report_background" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:31:37 | INFO | nipype.workflow | [Node] Finished
"ds_report_background", elapsed time 0.014363s.
2026-04-10 23:31:47 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_zoomed".
2026-04-10 23:31:47 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_zoomed" in "/home/jovyan/workspace/books/examples/wor
kflows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/ds_r
eport_zoomed".
2026-04-10 23:31:47 | INFO | nipype.workflow |34;2
0m [Node] Outdated cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_zoomed".
2026-04-10 23:31:47 | INFO | nipype.workflow | [Node] Executing
"ds_report_zoomed" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:31:47 | INFO | nipype.workflow | [Node] Finished
"ds_report_zoomed", elapsed time 0.009816s.
2026-04-10 23:31:47 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_bmask".
2026-04-10 23:31:47 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_bmask" in "/home/jovyan/workspace/books/examples/work
flows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/ds_re
port_bmask".
2026-04-10 23:31:47 | INFO | nipype.workflow |
[Node] Outdated cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_bmask".
2026-04-10 23:31:47 | INFO | nipype.workflow | [Node] Executing
"ds_report_bmask" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:31:47 | INFO | nipype.workflow | [Node] Finished
"ds_report_bmask", elapsed time 0.010172s.
2026-04-10 23:31:49 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_norm".
2026-04-10 23:31:49 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_norm" in "/home/jovyan/workspace/books/examples/workf
lows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/ds_rep
ort_norm".
2026-04-10 23:31:49 | INFO | nipype.workflow | [
Node] Outdated cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_norm".
2026-04-10 23:31:49 | INFO | nipype.workflow | [Node] Executing
"ds_report_norm" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:31:49 | INFO | nipype.workflow | [Node] Finished
"ds_report_norm", elapsed time 0.011417s.
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_mean".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_mean".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_mean0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/func
MRIQC/func_report_wf/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/ds_report_mean/mapflow/_ds_report_mean
0".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_mean0".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Executing
"_ds_report_mean0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Finished
"_ds_report_mean0", elapsed time 0.010794s.
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_background".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_background".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_background0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_w
f/funcMRIQC/func_report_wf/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/ds_report_background/mapflow/_ds
_report_background0".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_background0".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Executing
"_ds_report_background0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Finished
"_ds_report_background0", elapsed time 0.009354s.
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_stdev".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_stdev".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_stdev0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/fun
cMRIQC/func_report_wf/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/ds_report_stdev/mapflow/_ds_report_st
dev0".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_stdev0".
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Executing
"_ds_report_stdev0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:31:55 | INFO | nipype.workflow | [Node] Finished
"_ds_report_stdev0", elapsed time 0.011876s.
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_fa".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_fa" in "/home/jovyan/workspace/books/examples/workflows
/MRIQC_workdir/mriqc_wf/dwiMRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_fa
".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Ou
tdated cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_fa".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Executing
"ds_report_fa" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Finished
"ds_report_fa", elapsed time 0.012075s.
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_md".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_md" in "/home/jovyan/workspace/books/examples/workflows
/MRIQC_workdir/mriqc_wf/dwiMRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_md
".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Ou
tdated cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_md".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Executing
"ds_report_md" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Finished
"ds_report_md", elapsed time 0.010034s.
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_hm".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_hm" in "/home/jovyan/workspace/books/examples/workflows
/MRIQC_workdir/mriqc_wf/dwiMRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_hm
".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Ou
tdated cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_hm".
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Executing
"ds_report_hm" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:03 | INFO | nipype.workflow | [Node] Finished
"ds_report_hm", elapsed time 0.011322s.
2026-04-10 23:32:05 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_norm".
2026-04-10 23:32:05 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.funcMRIQC.func_report_wf.ds_report_norm" in "/home/jovyan/workspace/books/examples/workf
lows/MRIQC_workdir/mriqc_wf/funcMRIQC/func_report_wf/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/ds_rep
ort_norm".
2026-04-10 23:32:05 | INFO | nipype.workflow | [
Node] Outdated cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_norm".
2026-04-10 23:32:05 | INFO | nipype.workflow | [Node] Executing
"ds_report_norm" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:05 | INFO | nipype.workflow | [Node] Finished
"ds_report_norm", elapsed time 0.010666s.
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_zoomed".
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_zoomed".
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_zoomed0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/fu
ncMRIQC/func_report_wf/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/ds_report_zoomed/mapflow/_ds_report_
zoomed0".
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_zoomed0".
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Executing
"_ds_report_zoomed0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Finished
"_ds_report_zoomed0", elapsed time 0.01076s.
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_bmask".
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.funcMRIQC.func_report_wf.ds_report_bmask" in "/home/jovyan/workspace/books/examples/work
flows/MRIQC_workdir/mriqc_wf/funcMRIQC/func_report_wf/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/ds_re
port_bmask".
2026-04-10 23:32:07 | INFO | nipype.workflow |
[Node] Outdated cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_bmask".
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Executing
"ds_report_bmask" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Finished
"ds_report_bmask", elapsed time 0.011326s.
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Setting-u
p "_datasink0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/funcMRIQC/
ComputeIQMs/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/datasink/mapflow/_datasink0".
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Executing
"_datasink0" <mriqc.interfaces.bids.IQMFileSink>
2026-04-10 23:32:07 | INFO | nipype.workflow | [Node] Finished
"_datasink0", elapsed time 0.00167s.
2026-04-10 23:32:08 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_timeseries0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/funcM
RIQC/ComputeIQMs/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/ds_timeseries/mapflow/_ds_timeseries0".0
m
2026-04-10 23:32:08 | INFO | nipype.workflow | [Node] Executing
"_ds_timeseries0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:08 | INFO | nipype.workflow | [Node] Finished
"_ds_timeseries0", elapsed time 0.008834s.
2026-04-10 23:32:11 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_carpet".
2026-04-10 23:32:11 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.funcMRIQC.func_report_wf.ds_report_carpet".
2026-04-10 23:32:11 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_carpet0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/fu
ncMRIQC/func_report_wf/31cb705173c95b4fbebf2ff1b1a2240a80bc1460/ds_report_carpet/mapflow/_ds_report_
carpet0".
2026-04-10 23:32:11 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_carpet0".
2026-04-10 23:32:11 | INFO | nipype.workflow | [Node] Executing
"_ds_report_carpet0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:11 | INFO | nipype.workflow | [Node] Finished
"_ds_report_carpet0", elapsed time 0.010115s.
2026-04-10 23:32:15 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_headmask".
2026-04-10 23:32:15 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_headmask" in "/home/jovyan/workspace/books/examples/w
orkflows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/ds
_report_headmask".
2026-04-10 23:32:15 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_headmask".
2026-04-10 23:32:15 | INFO | nipype.workflow | [Node] Executing
"ds_report_headmask" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:15 | INFO | nipype.workflow | [Node] Finished
"ds_report_headmask", elapsed time 0.013025s.
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_segm".
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_segm" in "/home/jovyan/workspace/books/examples/workf
lows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/ds_rep
ort_segm".
2026-04-10 23:32:19 | INFO | nipype.workflow | [
Node] Outdated cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_segm".
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Executing
"ds_report_segm" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Finished
"ds_report_segm", elapsed time 0.009264s.
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_noisefit".
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_noisefit" in "/home/jovyan/workspace/books/examples/w
orkflows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/ds
_report_noisefit".
2026-04-10 23:32:19 | INFO | nipype.workflow |
34;20m [Node] Outdated cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_noisefit".
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Executing
"ds_report_noisefit" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Finished
"ds_report_noisefit", elapsed time 0.009007s.
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_airmask".
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_airmask" in "/home/jovyan/workspace/books/examples/wo
rkflows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/ds_
report_airmask".
2026-04-10 23:32:19 | INFO | nipype.workflow |34
;20m [Node] Outdated cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_airmask".
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Executing
"ds_report_airmask" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Finished
"ds_report_airmask", elapsed time 0.010058s.
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_artmask".
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_artmask" in "/home/jovyan/workspace/books/examples/wo
rkflows/MRIQC_workdir/mriqc_wf/anatMRIQC/anat_report_wf/867c4dee1c01330cc041483ba84642cbd1c2d8b1/ds_
report_artmask".
2026-04-10 23:32:19 | INFO | nipype.workflow |34
;20m [Node] Outdated cache found for "mriqc_wf.anatMRIQC.anat_report_wf.ds_report_artmask".
2026-04-10 23:32:19 | INFO | nipype.workflow | [Node] Executing
"ds_report_artmask" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:20 | INFO | nipype.workflow | [Node] Finished
"ds_report_artmask", elapsed time 0.011923s.
2026-04-10 23:32:20 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.anatMRIQC.ComputeIQMs.datasink" in "/home/jovyan/workspace/books/examples/workflows/MRIQ
C_workdir/mriqc_wf/anatMRIQC/ComputeIQMs/867c4dee1c01330cc041483ba84642cbd1c2d8b1/datasink".
2026-04-10 23:32:20 | INFO | nipype.workflow | [Node] Executing
"datasink" <mriqc.interfaces.bids.IQMFileSink>
2026-04-10 23:32:20 | INFO | nipype.workflow | [Node] Finished
"datasink", elapsed time 0.001213s.
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_noise0".
2026-04-10 23:32:33 | INFO | nipype
.workflow | [Node] Setting-up "_ds_report_noise0" in "/home/jovyan/workspace/books/exam
ples/workflows/MRIQC_workdir/mriqc_wf/dwiMRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb5
0/ds_report_noise/mapflow/_ds_report_noise0".
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_noise0".
2026-04-10 23:32:33 | INFO | nipype
.workflow | [Node] Executing "_ds_report_noise0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Finished
"_ds_report_noise0", elapsed time 0.009659s.
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_noise1".
2026-04-10 23:32:33 | INFO | nipype
.workflow | [Node] Setting-up "_ds_report_noise1" in "/home/jovyan/workspace/books/exam
ples/workflows/MRIQC_workdir/mriqc_wf/dwiMRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb5
0/ds_report_noise/mapflow/_ds_report_noise1".
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_noise1".
2026-04-10 23:32:33 | INFO | nipype
.workflow | [Node] Executing "_ds_report_noise1" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Finished
"_ds_report_noise1", elapsed time 0.010566s.
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_noise2".
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_noise2" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwi
MRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_noise/mapflow/_ds_report_nois
e2".
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_noise2".
2026-04-10 23:32:33 | INFO | nipype
.workflow | [Node] Executing "_ds_report_noise2" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Finished
"_ds_report_noise2", elapsed time 0.009362s.
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_noise3".
2026-04-10 23:32:33 | INFO | nipype
.workflow | [Node] Setting-up "_ds_report_noise3" in "/home/jovyan/workspace/books/exam
ples/workflows/MRIQC_workdir/mriqc_wf/dwiMRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb5
0/ds_report_noise/mapflow/_ds_report_noise3".
2026-04-10 23:32:33 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_noise3".
2026-04-10 23:32:34 | INFO | nipype
.workflow | [Node] Executing "_ds_report_noise3" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:34 | INFO | nipype.workflow | [Node] Finished
"_ds_report_noise3", elapsed time 0.008339s.
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_noise".
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_noise".
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_noise0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwi
MRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_noise/mapflow/_ds_report_nois
e0".
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Executing
"_ds_report_noise0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Finished
"_ds_report_noise0", elapsed time 0.009978s.
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_noise1" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwi
MRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_noise/mapflow/_ds_report_nois
e1".
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node]
Executing "_ds_report_noise1" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Finished
"_ds_report_noise1", elapsed time 0.010542s.
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_noise2" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwi
MRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_noise/mapflow/_ds_report_nois
e2".
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Executing
"_ds_report_noise2" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Finished
"_ds_report_noise2", elapsed time 0.009922s.
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_noise3" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwi
MRIQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_noise/mapflow/_ds_report_nois
e3".
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Executing
"_ds_report_noise3" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Finished
"_ds_report_noise3", elapsed time 0.009175s.
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Setting-u
p "mriqc_wf.dwiMRIQC.ComputeIQMs.datasink" in "/home/jovyan/workspace/books/examples/workflows/MRIQC
_workdir/mriqc_wf/dwiMRIQC/ComputeIQMs/edbb42c7bf5828aa552d68da572c7aebc01fdb50/datasink".
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Executing
"datasink" <mriqc.interfaces.bids.IQMFileSink>
2026-04-10 23:32:35 | INFO | nipype.workflow | [Node] Finished
"datasink", elapsed time 0.001355s.
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_snr0".
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_snr0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwiMR
IQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_snr/mapflow/_ds_report_snr0".
0m
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Outdat
ed cache found for "_ds_report_snr0".
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Executing
"_ds_report_snr0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Finished
"_ds_report_snr0", elapsed time 0.011876s.
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_snr1".
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_snr1" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwiMR
IQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_snr/mapflow/_ds_report_snr1".
0m
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Outdat
ed cache found for "_ds_report_snr1".
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Executing
"_ds_report_snr1" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Finished
"_ds_report_snr1", elapsed time 0.01011s.
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_snr2".
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_snr2" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwiMR
IQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_snr/mapflow/_ds_report_snr2".
0m
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_snr2".
2026-04-10 23:32:41 | INFO | nipype.w
orkflow | [Node] Executing "_ds_report_snr2" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Finished
"_ds_report_snr2", elapsed time 0.010052s.
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_snr3".
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_snr3" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwiMR
IQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_snr/mapflow/_ds_report_snr3".
0m
2026-04-10 23:32:41 | INFO | nipype.workflow | [Node] Outdated
cache found for "_ds_report_snr3".
2026-04-10 23:32:41 | INFO | nipype.w
orkflow | [Node] Executing "_ds_report_snr3" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:42 | INFO | nipype.workflow | [Node] Finished
"_ds_report_snr3", elapsed time 0.009546s.
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_snr".
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Outdated
cache found for "mriqc_wf.dwiMRIQC.dwi_report_wf.ds_report_snr".
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_snr0" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwiMR
IQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_snr/mapflow/_ds_report_snr0".
0m
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Executing
"_ds_report_snr0" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Finished
"_ds_report_snr0", elapsed time 0.011305s.
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_snr1" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwiMR
IQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_snr/mapflow/_ds_report_snr1".
0m
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Executing
"_ds_report_snr1" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Finished
"_ds_report_snr1", elapsed time 0.010464s.
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_snr2" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwiMR
IQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_snr/mapflow/_ds_report_snr2".
0m
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Executing
"_ds_report_snr2" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Finished
"_ds_report_snr2", elapsed time 0.009727s.
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Setting-u
p "_ds_report_snr3" in "/home/jovyan/workspace/books/examples/workflows/MRIQC_workdir/mriqc_wf/dwiMR
IQC/dwi_report_wf/edbb42c7bf5828aa552d68da572c7aebc01fdb50/ds_report_snr/mapflow/_ds_report_snr3".
0m
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Executing
"_ds_report_snr3" <mriqc.interfaces.DerivativesDataSink>
2026-04-10 23:32:43 | INFO | nipype.workflow | [Node] Finished
"_ds_report_snr3", elapsed time 0.010577s.
2026-04-10 23:32:45 | INFO | nipype.workflow | Generating repor
ts...
2026-04-10 23:32:46 | INFO | nipype.workflow | Report generatio
n finished (3 reports).
2026-04-10 23:32:46 | IMPORTANT | mriqc
[0m| Participant level finished successfully averaging 00h 00min 34s per subject.
2026-04-10 23:32:46 | INFO | mriqc | Generating BIDS
derivatives metadata.
----------------------------------------------------------------
MRIQC c
ompleted (elapsed time 00h 01min 44s).
-------------------------------------------------------------
---
The carpet plot offers a compact way to visualize voxel-wise fMRI signal changes over time, allowing quick identification of artifacts, motion spikes, or other irregularities in the data. Each row corresponds to a voxel (or a group of voxels), and each column represents a time point, effectively summarizing the entire 4D dataset in a single image.
Here’s an example of a carpet plot generated by MRIQC for subject sub-CON02 during the resting-state session:
SVG(filename='./MRIQC/sub-CON02/figures/sub-CON02_ses-preop_task-rest_desc-carpet_bold.svg')
4.2 Core Neuroimaging Tools (Command-Line Interfaces)#
The following tools are widely used in neuroimaging research and are typically run from the command line. They support key tasks in structural, functional, and diffusion imaging workflows. Many are integrated into higher-level toolkits like Nipype or fMRIPrep.
# create an output directory
!mkdir -p output
FreeSurfer’s SynthStrip: Brain extraction of the anatomical image#
Another interactive example can be found here: recon-all
# SynthStrip
! mri_synthstrip -i ./ds001226/sub-CON02/ses-preop/anat/sub-CON02_ses-preop_T1w.nii.gz -o ./output/synth_stripped.nii.gz -m ./output/synth_mask.nii.gz
Configuring model on the CPU
Running SynthStrip model version 1
Input image read from: ./ds001226/sub-CON02/ses-preop/anat/sub-CON02_ses-preop_T1w.nii.gz
Processing frame (of 1): 1
done
Masked image saved to: ./output/synth_stripped.nii.gz
Binary brain mask saved to: ./output/synth_mask.nii.gz
If you use SynthStrip in your analysis, please cite:
----------------------------------------------------
SynthStrip: Skull-Stripping for Any Brain Image
A Hoopes, JS Mora, AV Dalca, B Fischl, M Hoffmann
NeuroImage 206 (2022), 119474
https://doi.org/10.1016/j.neuroimage.2022.119474
Website: https://synthstrip.io
AFNI’s 3dvolreg: Motion Correction of functional data#
For a complete example about preprocessing and glm, have a look at this AFNI Preprocessing and GLM notebook.
# 3dvolreg
! 3dvolreg -verbose -zpad 1 -cubic -base 0 -prefix ./output/epi_volreg.nii.gz -1Dfile ./output/motion.1D ./ds001226/sub-CON02/ses-preop/func/sub-CON02_ses-preop_task-rest_bold.nii.gz
++ 3dvolreg: AFNI version=AFNI_21.2.00 (Jul 8 2021) [64-bit]
++ Authored by: RW Cox
*+ WARNING: If you are performing spatial transformations on an oblique dset,
such as ./ds001226/sub-CON02/ses-preop/func/sub-CON02_ses-preop_task-rest_bold.nii.gz,
or viewing/combining it with volumes of differing obliquity,
you should consider running:
3dWarp -deoblique
on this and other oblique datasets in the same session.
See 3dWarp -help for details.
++ Oblique dataset:./ds001226/sub-CON02/ses-preop/func/sub-CON02_ses-preop_task-rest_bold.nii.gz is 8.917687 degrees from plumb.
++ Reading input dataset ./ds001226/sub-CON02/ses-preop/func/sub-CON02_ses-preop_task-rest_bold.nii.gz
++ Edging: x=3 y=3 z=2
++ Creating mask for -maxdisp
+ Automask has 50248 voxels
+ 6670 voxels left in -maxdisp mask after erosion
++ Initializing alignment base
++ Starting final pass on 180 sub-bricks: 0..1..2
..3
..4
..5
..6
..7
..8
..9
..10
..11
..12
..13
..14
..15
..16
..17
..18
..19..20
..21
..22
..23
..24
..25
..26
..27
..28..29
..30
..31
..32
..33
..34
..35
..36
..37
..38
..39
..40..41
..42
..43
..44
..45
..46
..47
..48
..49
..50
..51
.
.52..53
..54
..55
..56
..57
..58
..59
..60
..61..62
..63
..64
..65
..66
..67
..68
..69
..70
..71
..72
..73
..74
..75
..76
..77
..78
..79
..80
..81
..82
..83
..84
..85
..86
..87
..88
..89
..90
..91
..92
..93
..94
..95
..96
..97
..98
..99
..100
..101
..102
..103
..104
..105
..106
..107
..108
..109
..110
..111
..112
..113
..114
..115
..116
..117
..118
..119
..120
..121
..122
..123
..124
..125
..126
.
.127
..128
..129
..130
..131
..132
..133
..134
..135
..136
..137
..138
..139
..140
..141
..142
..143
..144
..145
..146
..147
..148
..149
..150
..151
..152
..153
..154
..155
..156
..157
..158
..159
..160
..161
..162
..163
..164
..165
..166
.
.167
..168
..169
..170
..171
..172
..173
..174
..175
..176
..177
..178
..179
..
++ CPU time for realignment=0 s [=0 s/sub-brick]
++ Min : roll=-0.392 pitch=-0.667 yaw=-0.087 dS=-1.333 dL=-0.196 dP=-0.715
++ Mean: roll=-0.183 pitch=-0.191 yaw=+0.040 dS=-0.527 dL=-0.064 dP=-0.354
++ Max : roll=+0.016 pitch=+0.160 yaw=+0.166 dS=+0.086 dL=+0.149 dP=+0.183
++ Max displacements (mm) for each sub-brick:
0.00(0.00) 0.44(0.44) 0.27(0.39) 0.12(0.23) 0.44(0.37) 0.47(0.19) 0.21(0.45) 0.42(0.42) 0.48(0.15) 0.26(0.40) 0.38(0.50) 0.52(0.15) 0.28(0.49) 0.28(0.29) 0.55(0.34) 0.28(0.45) 0.28(0.19) 0.48(0.36) 0.56(0.13) 0.35(0.38) 0.34(0.27) 0.53(0.29) 0.45(0.38) 0.38(0.20) 0.52(0.35) 0.57(0.16) 0.58(0.11) 0.60(0.27) 0.44(0.33) 0.54(0.36) 0.58(0.30) 0.42(0.39) 0.50(0.31) 0.64(0.15) 0.42(0.42) 0.45(0.35) 0.69(0.30) 0.56(0.47) 0.68(0.48) 0.75(0.26) 0.48(0.55) 0.60(0.42) 0.81(0.37) 0.59(0.53) 0.68(0.37) 0.55(0.39) 0.67(0.29) 0.82(0.39) 0.61(0.41) 0.75(0.29) 0.58(0.37) 0.60(0.22) 0.60(0.25) 0.49(0.47) 0.46(0.32) 0.57(0.26) 0.52(0.40) 0.57(0.35) 0.55(0.35) 0.57(0.32) 0.64(0.29) 0.53(0.41) 0.69(0.55) 0.59(0.41) 0.60(0.25) 0.91(0.64) 0.71(0.59) 0.68(0.28) 0.69(0.34) 0.71(0.31) 0.73(0.26) 0.88(0.51) 0.71(0.45) 1.09(0.64) 1.04(0.35) 1.13(0.14) 0.93(0.46) 0.95(0.28) 0.75(0.29) 0.92(0.27) 0.91(0.42) 0.74(0.41) 1.03(0.43) 0.82(0.26) 1.17(0.41) 0.91(0.32) 1.12(0.36) 1.06(0.36) 1.01(0.43) 1.67(0.96) 1.10(0.76) 1.72(0.76) 1.45(0.33) 1.22(0.59) 1.28(0.32) 1.08(0.41) 1.28(0.36) 1.06(0.31) 1.60(0.67) 1.38(0.46) 1.48(0.54) 1.69(0.48) 1.77(0.55) 1.99(0.49) 1.53(0.56) 1.59(0.23) 1.51(0.45) 1.46(0.57) 1.57(0.37) 1.36(0.36) 1.60(0.30) 1.37(0.26) 1.73(0.42) 1.36(0.47) 2.12(0.90) 1.64(0.59) 2.12(0.57) 1.58(0.68) 2.11(0.69) 1.79(0.42) 1.69(0.38) 1.64(0.34) 1.49(0.21) 1.56(0.38) 1.61(0.22) 1.55(0.28) 1.49(0.38) 1.66(0.30) 1.45(0.24) 1.50(0.37) 1.50(0.19) 1.42(0.12) 1.38(0.26) 1.48(0.16) 1.32(0.22) 1.26(0.28) 1.37(0.13) 1.27(0.32) 1.17(0.20) 1.32(0.32) 1.29(0.25) 1.18(0.14) 1.29(0.33) 1.34(0.24) 1.26(0.16) 1.22(0.30) 1.33(0.19) 1.31(0.24) 1.21(0.32) 1.55(0.45) 1.45(0.43) 1.52(0.55) 2.03(0.70) 1.66(0.52) 1.47(0.30) 1.48(0.36) 1.50(0.34) 1.33(0.30) 1.64(0.38) 1.43(0.28) 1.64(0.26) 1.35(0.38) 1.61(0.30) 1.48(0.29) 1.55(0.44) 1.47(0.46) 1.46(0.46) 1.57(0.35) 1.46(0.26) 2.09(0.82) 1.69(0.57) 1.94(0.43) 1.79(0.43) 1.66(0.24) 2.27(0.88) 1.94(0.59) 2.26(0.98) 2.14(0.93) 1.75(0.51) 1.78(0.31)
++ Max displacement in automask = 2.27 (mm) at sub-brick 174
++ Max delta displ in automask = 0.98 (mm) at sub-brick 176
** ERROR: output dataset name 'epi_volreg.nii.gz' conflicts with existing file
** ERROR: dataset NOT written to disk!
++ Wrote dataset to disk in ./output/epi_volreg_AA1.nii.gz
** Warning: overwriting file ./output/motion.1D
FSL: Registration of the anatomical image to MNI#
A detailed example notebook about FSL Preprocessing and GLM is also available.
fsl_path = !which fsl
fsl_version = fsl_path[0].split('_')[1]
MNI_BRAIN = os.path.join(
'/opt',
f'fsl-{fsl_version}',
'data',
'standard',
'MNI152_T1_2mm_brain.nii.gz'
)
print(MNI_BRAIN)
/opt/fsl-6.0.7.16/data/standard/MNI152_T1_2mm_brain.nii.gz
# Register the T1w image to the MNI template (default: dof 12, normal search)
!flirt -in ./output/synth_stripped.nii.gz -ref "{MNI_BRAIN}" -out ./output/synth_stripped_flirt_T1MNI.nii.gz -omat ./output/synth_stripped_flirt_T1MNI.mat
MRTrix: Denoising diffusion data#
There is a series of 3 notebooks about MRTrix: Part 1 - Preprocessing can be found here.
# converts diffusion data to a format that MRtrix understands
! mrconvert ./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz ./output/sub-02_dwi.mif -fslgrad ./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.bvec ./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.bval -force
mrconvert: [WARNING] existing output files will be overwritten
mrconvert: [ 0%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 4%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 8%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 13%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 17%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 21%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 26%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 31%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 36%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 41%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 46%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 51%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 56%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 62%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 67%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 73%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 78%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 84%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 91%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [ 98%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"...?7h
mrconvert: [100%] uncompressing image "./ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz"
mrconvert: [ 36%] copying from "./ds001226...ses-preop_acq-AP_dwi.nii.gz" to "./output/sub-02_dwi.mif"...?7h
mrconvert: [ 73%] copying from "./ds001226...ses-preop_acq-AP_dwi.nii.gz" to "./output/sub-02_dwi.mif"...?7h
mrconvert: [100%] copying from "./ds001226...ses-preop_acq-AP_dwi.nii.gz" to "./output/sub-02_dwi.mif"
# denoising
! dwidenoise ./output/sub-02_dwi.mif ./output/sub-02_den.mif -noise ./output/noise.mif -force
dwidenoise: [WARNING] existing output files will be overwritten
dwidenoise: [ 1%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 9%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 17%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 25%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 33%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 42%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 50%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 59%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 67%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 75%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 84%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [ 92%] preloading data for "./output/sub-02_dwi.mif"...?7h
dwidenoise: [100%] preloading data for "./output/sub-02_dwi.mif"
dwidenoise: [ 0%] running MP-PCA denoising...?7h
dwidenoise: [ 1%] running MP-PCA denoising...?7h
dwidenoise: [ 2%] running MP-PCA denoising...?7h
dwidenoise: [ 3%] running MP-PCA denoising...?7h
dwidenoise: [ 4%] running MP-PCA denoising...?7h
dwidenoise: [ 5%] running MP-PCA denoising...?7h
dwidenoise: [ 6%] running MP-PCA denoising...?7h
dwidenoise: [ 7%] running MP-PCA denoising...?7h
dwidenoise: [ 8%] running MP-PCA denoising...?7h
dwidenoise: [ 9%] running MP-PCA denoising...?7h
dwidenoise: [ 10%] running MP-PCA denoising...?7h
dwidenoise: [ 11%] running MP-PCA denoising...?7h
dwidenoise: [ 12%] running MP-PCA denoising...?7h
dwidenoise: [ 13%] running MP-PCA denoising...?7h
dwidenoise: [ 14%] running MP-PCA denoising...?7h
dwidenoise: [ 15%] running MP-PCA denoising...?7h
dwidenoise: [ 16%] running MP-PCA denoising...?7h
dwidenoise: [ 17%] running MP-PCA denoising...?7h
dwidenoise: [ 18%] running MP-PCA denoising...?7h
dwidenoise: [ 19%] running MP-PCA denoising...?7h
dwidenoise: [ 20%] running MP-PCA denoising...?7h
dwidenoise: [ 21%] running MP-PCA denoising...?7h
dwidenoise: [ 22%] running MP-PCA denoising...?7h
dwidenoise: [ 23%] running MP-PCA denoising...?7h
dwidenoise: [ 24%] running MP-PCA denoising...?7h
dwidenoise: [ 25%] running MP-PCA denoising...?7h
dwidenoise: [ 26%] running MP-PCA denoising...?7h
dwidenoise: [ 27%] running MP-PCA denoising...?7h
dwidenoise: [ 28%] running MP-PCA denoising...?7h
dwidenoise: [ 29%] running MP-PCA denoising...?7h
dwidenoise: [ 30%] running MP-PCA denoising...?7h
dwidenoise: [ 31%] running MP-PCA denoising...?7h
dwidenoise: [ 32%] running MP-PCA denoising...?7h
dwidenoise: [ 33%] running MP-PCA denoising...?7h
dwidenoise: [ 34%] running MP-PCA denoising...?7h
dwidenoise: [ 35%] running MP-PCA denoising...?7h
dwidenoise: [ 36%] running MP-PCA denoising...?7h
dwidenoise: [ 37%] running MP-PCA denoising...?7h
dwidenoise: [ 38%] running MP-PCA denoising...?7h
dwidenoise: [ 39%] running MP-PCA denoising...?7h
dwidenoise: [ 40%] running MP-PCA denoising...?7h
dwidenoise: [ 41%] running MP-PCA denoising...?7h
dwidenoise: [ 42%] running MP-PCA denoising...?7h
dwidenoise: [ 43%] running MP-PCA denoising...?7h
dwidenoise: [ 44%] running MP-PCA denoising...?7h
dwidenoise: [ 45%] running MP-PCA denoising...?7h
dwidenoise: [ 46%] running MP-PCA denoising...?7h
dwidenoise: [ 47%] running MP-PCA denoising...?7h
dwidenoise: [ 48%] running MP-PCA denoising...?7h
dwidenoise: [ 49%] running MP-PCA denoising...?7h
dwidenoise: [ 50%] running MP-PCA denoising...?7h
dwidenoise: [ 51%] running MP-PCA denoising...?7h
dwidenoise: [ 52%] running MP-PCA denoising...?7h
dwidenoise: [ 53%] running MP-PCA denoising...?7h
dwidenoise: [ 54%] running MP-PCA denoising...?7h
dwidenoise: [ 55%] running MP-PCA denoising...?7h
dwidenoise: [ 56%] running MP-PCA denoising...?7h
dwidenoise: [ 57%] running MP-PCA denoising...?7h
dwidenoise: [ 58%] running MP-PCA denoising...?7h
dwidenoise: [ 59%] running MP-PCA denoising...?7h
dwidenoise: [ 60%] running MP-PCA denoising...?7h
dwidenoise: [ 61%] running MP-PCA denoising...?7h
dwidenoise: [ 62%] running MP-PCA denoising...?7h
dwidenoise: [ 63%] running MP-PCA denoising...?7h
dwidenoise: [ 64%] running MP-PCA denoising...?7h
dwidenoise: [ 65%] running MP-PCA denoising...?7h
dwidenoise: [ 66%] running MP-PCA denoising...?7h
dwidenoise: [ 67%] running MP-PCA denoising...?7h
dwidenoise: [ 68%] running MP-PCA denoising...?7h
dwidenoise: [ 69%] running MP-PCA denoising...?7h
dwidenoise: [ 70%] running MP-PCA denoising...?7h
dwidenoise: [ 71%] running MP-PCA denoising...?7h
dwidenoise: [ 72%] running MP-PCA denoising...?7h
dwidenoise: [ 73%] running MP-PCA denoising...?7h
dwidenoise: [ 74%] running MP-PCA denoising...?7h
dwidenoise: [ 75%] running MP-PCA denoising...?7h
dwidenoise: [ 76%] running MP-PCA denoising...?7h
dwidenoise: [ 77%] running MP-PCA denoising...?7h
dwidenoise: [ 78%] running MP-PCA denoising...?7h
dwidenoise: [ 79%] running MP-PCA denoising...?7h
dwidenoise: [ 80%] running MP-PCA denoising...?7h
dwidenoise: [ 81%] running MP-PCA denoising...?7h
dwidenoise: [ 82%] running MP-PCA denoising...?7h
dwidenoise: [ 83%] running MP-PCA denoising...?7h
dwidenoise: [ 84%] running MP-PCA denoising...?7h
dwidenoise: [ 85%] running MP-PCA denoising...?7h
dwidenoise: [ 86%] running MP-PCA denoising...?7h
dwidenoise: [ 87%] running MP-PCA denoising...?7h
dwidenoise: [ 88%] running MP-PCA denoising...?7h
dwidenoise: [ 89%] running MP-PCA denoising...?7h
dwidenoise: [ 90%] running MP-PCA denoising...?7h
dwidenoise: [ 91%] running MP-PCA denoising...?7h
dwidenoise: [ 92%] running MP-PCA denoising...?7h
dwidenoise: [ 93%] running MP-PCA denoising...?7h
dwidenoise: [ 94%] running MP-PCA denoising...?7h
dwidenoise: [ 95%] running MP-PCA denoising...?7h
dwidenoise: [ 96%] running MP-PCA denoising...?7h
dwidenoise: [ 97%] running MP-PCA denoising...?7h
dwidenoise: [ 98%] running MP-PCA denoising...?7h
dwidenoise: [ 99%] running MP-PCA denoising...?7h
dwidenoise: [100%] running MP-PCA denoising...?7h
dwidenoise: [100%] running MP-PCA denoising
4.3 Pythonic Interfaces and Python Toolkits for Neuroimaging#
Nipype#
Nipype is a flexible Python-based workflow engine that enables users to build reproducible, modular pipelines by integrating diverse neuroimaging tools under a consistent interface. We will explore how it wraps individual tools such as fsl.BET() and afni.Edge3() directly from Python code.
An overview of Nipype and how to build pipelines can be found in Nipype on Neurodesk.
# this makes it so FSL knows how we want our output files
os.environ["FSLOUTPUTTYPE"]="NIFTI_GZ"
btr = fsl.BET()
btr.inputs.in_file = './ds001226/sub-CON02/ses-preop/anat/sub-CON02_ses-preop_T1w.nii.gz'
btr.inputs.frac = 0.4
btr.inputs.out_file = './output/nipype_bet.nii.gz'
res = btr.run()
edge3 = afni.Edge3()
edge3.inputs.in_file = './ds001226/sub-CON02/ses-preop/anat/sub-CON02_ses-preop_T1w.nii.gz'
edge3.inputs.out_file = './output/nipype_afni_edges.nii.gz'
edge3.inputs.datum = 'byte'
res = edge3.run()
260410-23:34:26,102 nipype.interface INFO:
stderr 2026-04-10T23:34:26.102445:++ 3dedge3: AFNI version=AFNI_21.2.00 (Jul 8 2021) [64-bit]
260410-23:34:26,108 nipype.interface INFO:
stderr 2026-04-10T23:34:26.108525:*+ WARNING: If you are performing spatial transformations on an oblique dset,
260410-23:34:26,109 nipype.interface INFO:
stderr 2026-04-10T23:34:26.108525: such as ds001226/sub-CON02/ses-preop/anat/sub-CON02_ses-preop_T1w.nii.gz,
260410-23:34:26,110 nipype.interface INFO:
stderr 2026-04-10T23:34:26.108525: or viewing/combining it with volumes of differing obliquity,
260410-23:34:26,111 nipype.interface INFO:
stderr 2026-04-10T23:34:26.108525: you should consider running:
260410-23:34:26,112 nipype.interface INFO:
stderr 2026-04-10T23:34:26.108525: 3dWarp -deoblique
260410-23:34:26,113 nipype.interface INFO:
stderr 2026-04-10T23:34:26.108525: on this and other oblique datasets in the same session.
260410-23:34:26,114 nipype.interface INFO:
stderr 2026-04-10T23:34:26.108525: See 3dWarp -help for details.
260410-23:34:26,115 nipype.interface INFO:
stderr 2026-04-10T23:34:26.108525:++ Oblique dataset:ds001226/sub-CON02/ses-preop/anat/sub-CON02_ses-preop_T1w.nii.gz is 3.067714 degrees from plumb.
260410-23:34:29,341 nipype.interface INFO:
stderr 2026-04-10T23:34:29.341779:** ERROR: output dataset name 'nipype_afni_edges.nii.gz' conflicts with existing file
260410-23:34:29,343 nipype.interface INFO:
stderr 2026-04-10T23:34:29.341779:** ERROR: dataset NOT written to disk!
ANTsPy#
ANTsPy is a Python wrapper for the widely-used ANTs (Advanced Normalization Tools) C++ biomedical image processing library. It offers fast reading and writing of medical images, advanced algorithms for image registration, segmentation, and statistical learning, as well as tools for generating publication-quality visualizations.
Segmentation#
The segmentation module provides methods such as Atropos segmentation, Joint Label Fusion, cortical thickness estimation, and prior-based segmentation.
Atropos segmentation:
img = ants.image_read(ants.get_ants_data('r16'))
img = ants.resample_image(img, (64,64), 1, 0)
mask = ants.get_mask(img)
img_seg = ants.atropos(a=img, m='[0.2,1x1]', c='[2,0]',
i='kmeans[3]', x=mask)
print(img_seg.keys())
ants.plot(img_seg['segmentation'])
dict_keys(['segmentation', 'probabilityimages'])
Cortical thickness
img = ants.image_read(ants.get_ants_data('r16'),2)
mask = ants.get_mask(img).threshold_image(1, 2)
segs = ants.atropos(a=img, m='[0.2,1x1]', c='[2,0]', i='kmeans[3]', x= mask)
thickimg = ants.kelly_kapowski(s=segs['segmentation'], g=segs['probabilityimages'][1],
w=segs['probabilityimages'][2], its=45,
r=0.5, m=1)
print(thickimg)
img.plot(overlay=thickimg, overlay_cmap='jet')
ANTsImage
Pixel Type : float (float32)
Components : 1
Dimensions : (256, 256)
Spacing : (1.0, 1.0)
Origin : (0.0, 0.0)
Direction : [1. 0. 0. 1.]
Registration#
The registration module features the core ANTs registration interface, enabling access to all registration algorithms. It also includes utilities for evaluating registration quality, resampling and reorienting images, and applying specific transformations.
Symmetric Normalization (SyN)
fixed = ants.image_read(ants.get_ants_data('r16')).resample_image((64,64),1,0)
moving = ants.image_read(ants.get_ants_data('r64')).resample_image((64,64),1,0)
fixed.plot(overlay=moving, title='Before Registration')
mytx = ants.registration(fixed=fixed , moving=moving, type_of_transform='SyN' )
print(mytx)
warped_moving = mytx['warpedmovout']
fixed.plot(overlay=warped_moving,title='After Registration')
{'warpedmovout': ANTsImage
Pixel Type : float (float32)
Components : 1
Dimensions : (64, 64)
Spacing : (4.0476, 4.0476)
Origin : (0.0, 0.0)
Direction : [1. 0. 0. 1.]
, 'warpedfixout': ANTsImage
Pixel Type : float (float32)
Components : 1
Dimensions : (64, 64)
Spacing : (4.0476, 4.0476)
Origin : (0.0, 0.0)
Direction : [1. 0. 0. 1.]
, 'fwdtransforms': ['/tmp/tmpdlx0xp381Warp.nii.gz', '/tmp/tmpdlx0xp380GenericAffine.mat'], 'invtransforms': ['/tmp/tmpdlx0xp380GenericAffine.mat', '/tmp/tmpdlx0xp381InverseWarp.nii.gz']}
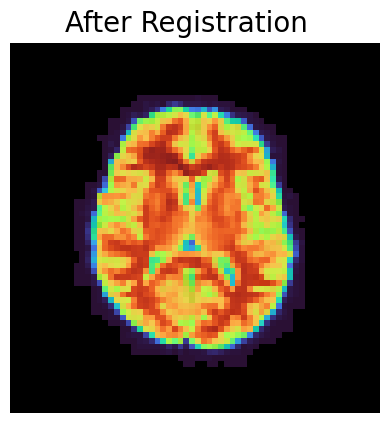
DIPY - Reconstruction of the diffusion signal with DTI (single tensor) model#
DIPY (Diffusion Imaging in Python) is a comprehensive library for processing and analyzing 3D and 4D medical imaging data. As part of the NiPy ecosystem, it provides robust tools for spatial normalization, denoising, tractography, and advanced diffusion modeling. In this section, we focus on reconstructing the diffusion signal using the classic Diffusion Tensor Imaging (DTI) model — a foundational approach for estimating white matter structure.
# Get data
fdwi = './ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.nii.gz'
fbval = './ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.bval'
fbvec = './ds001226/sub-CON02/ses-preop/dwi/sub-CON02_ses-preop_acq-AP_dwi.bvec'
data, affine, img = load_nifti(fdwi, return_img=True)
print(data.shape)
print(img.header.get_zooms()[:3])
(96, 96, 60, 102)
(np.float32(2.5), np.float32(2.5), np.float32(2.5))
# Load the b-values and b-vectors
bvals, bvecs = read_bvals_bvecs(fbval, fbvec)
gtab = gradient_table(bvals, bvecs=bvecs)
print(gtab.info)
B-values shape (102,)
min 0.000000
max 2800.000000
B-vectors shape (102, 3)
min -0.998927
max 0.997386
None
# Mask and crop the data
maskdata, mask = median_otsu(
data, vol_idx=range(10, 50), median_radius=3, numpass=1, autocrop=True, dilate=2
)
print(f"maskdata.shape {maskdata.shape}")
maskdata.shape (59, 73, 55, 102)
# Instantiate the Tensor model and fit the masked data
tenmodel = dti.TensorModel(gtab)
tenfit = tenmodel.fit(maskdata)
# Initialize a NIfTI header and set its intent to 'vector' to indicate multi-component data (e.g., DEC map)
hdr = nib.Nifti1Header()
hdr.set_intent('vector')
hdr['intent_name'] = b'RGB'
# Compute the color FA map (also known as the DEC map) from the diffusion tensor model,
# and save it as a NIfTI image with the appropriate vector metadata
img = nib.Nifti1Image(tenfit.color_fa, affine, header=hdr)
nib.save(img, './output/colorfa.nii.gz')
# Fit the tensor Orientation Distribution Functions on a subset of the data
sphere = get_sphere(name="repulsion724")
tensor_odfs = tenmodel.fit(data[25:60, 30:70, 29:30]).odf(sphere)
scene = window.Scene()
odf_actor = actor.odf_slicer(tensor_odfs, sphere=sphere, scale=0.5, colormap=None)
scene.add(odf_actor)
# Save a snapshot of the rendered scene as a PNG image to display it in a later step or to share it outside the notebook
window.record(scene=scene, n_frames=1, out_path="./output/tensor_odfs.png", size=(600, 600))
5. Visualization: The Art of Brain Mapping#
DIPY#
Visualizing Tensor Orientation Distributions with DIPY and IPython#
display(Image(filename='./output/tensor_odfs.png'))
Creating a mosaic plot#
t1_nipype = './output/nipype_bet.nii.gz'
data, affine = load_nifti(t1_nipype)
#Render slices from T1 with a specific value range
mean, std = data[data > 0].mean(), data[data > 0].std()
value_range = (mean - 0.5 * std, mean + 1.5 * std)
slice_actor = actor.slicer(data, affine=affine, value_range=value_range)
scene = window.Scene()
scene.clear()
scene.projection(proj_type="parallel")
cnt = 0
X, Y, Z = slice_actor.shape[:3]
rows, cols, border = 20, 15, 5
for j in range(rows):
for i in range(cols):
slice_mosaic = slice_actor.copy()
slice_mosaic.display(z=cnt)
slice_mosaic.SetPosition(
(X + border) * i, 0.5 * cols * (Y + border) - (Y + border) * j, 0
)
slice_mosaic.SetInterpolate(False)
scene.add(slice_mosaic)
cnt += 1
if cnt > Z:
break
if cnt > Z:
break
scene.reset_camera()
scene.zoom(3.5) #zoom in
window.record(scene=scene, out_path="./output/mosaic.png", size=(800, 800), reset_camera=False)
display(Image(filename='./output/mosaic.png'))

Matplotlib#
Brain slices#
# View 3D data
def view_slices_3d(image_3d, slice_nbr, vmin, vmax, title=''):
fig = plt.figure(figsize=(15, 4))
plt.suptitle(title, fontsize=14)
plt.subplot(131)
plt.imshow(np.take(image_3d, slice_nbr, 2), vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Axial', fontsize=10);
plt.subplot(132)
image_rot = ndimage.rotate(np.take(image_3d, slice_nbr, 1),90)
plt.imshow(image_rot, vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Coronal', fontsize=10);
plt.subplot(133)
image_rot = ndimage.rotate(np.take(image_3d, slice_nbr, 0),90)
plt.imshow(image_rot, vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Sagittal', fontsize=10);
cbar=plt.colorbar()
# load data
brain_full = nib.load('./ds001226/sub-CON02/ses-preop/anat/sub-CON02_ses-preop_T1w.nii.gz').get_fdata()
brain_edges = nib.load('./output/nipype_afni_edges.nii.gz').get_fdata()
view_slices_3d(brain_full, slice_nbr=135, vmin=0, vmax=500, title='Brain and Skull')
view_slices_3d(brain_edges, slice_nbr=135, vmin=0, vmax=500, title='Brain Edges')
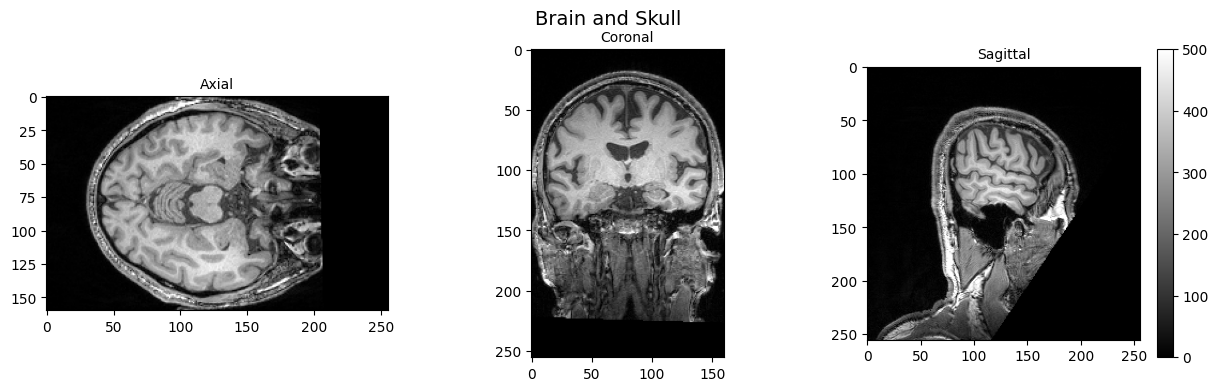
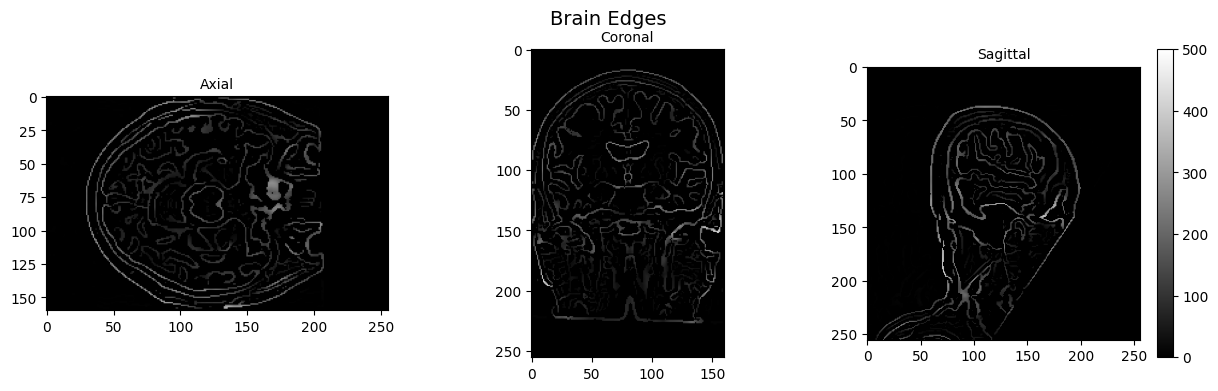
Motion parameters#
# Load AFNI motion parameters (roll, pitch, yaw, dS, dL, dP)
motion_params = np.loadtxt('./output/motion.1D')
timepoints = np.arange(motion_params.shape[0])
# Plotting
fig, axs = plt.subplots(2, 1, figsize=(12, 6), sharex=True)
# Rotations in degrees
axs[0].plot(timepoints, motion_params[:, 0], label='Roll (I-S)')
axs[0].plot(timepoints, motion_params[:, 1], label='Pitch (R-L)')
axs[0].plot(timepoints, motion_params[:, 2], label='Yaw (A-P)')
axs[0].set_ylabel('Rotation (degrees)')
axs[0].legend()
axs[0].set_title('AFNI Motion Parameters')
# Translations in mm
axs[1].plot(timepoints, motion_params[:, 3], label='dS (Superior)')
axs[1].plot(timepoints, motion_params[:, 4], label='dL (Left)')
axs[1].plot(timepoints, motion_params[:, 5], label='dP (Posterior)')
axs[1].set_ylabel('Translation (mm)')
axs[1].set_xlabel('Timepoint (volume)')
axs[1].legend()
plt.tight_layout()
plt.show()
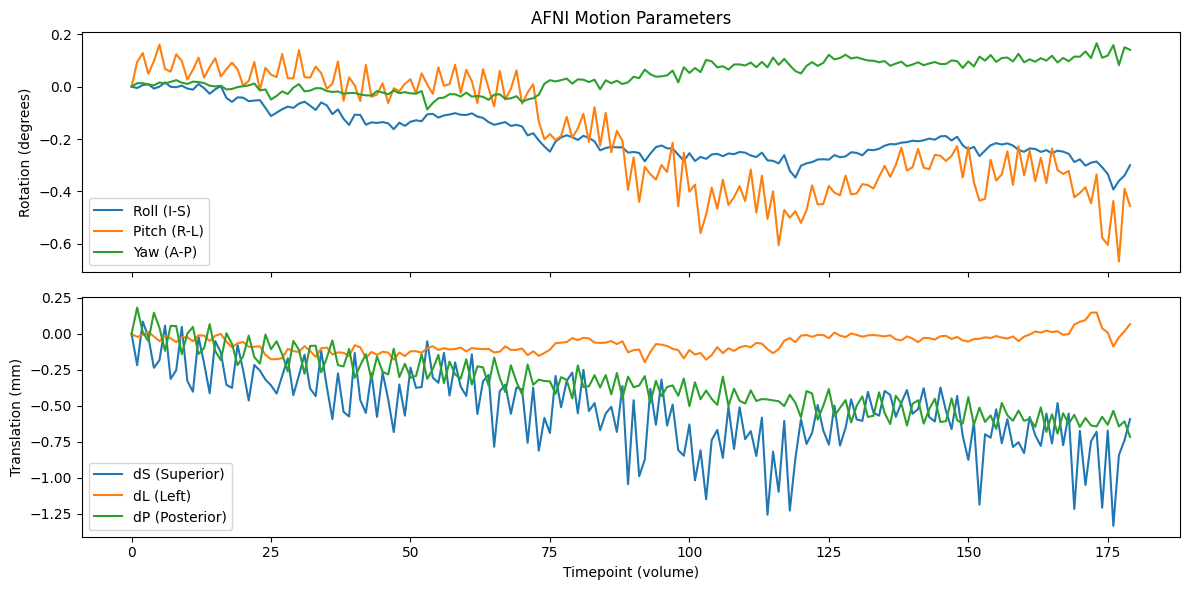
Nilearn#
plot_epi#
Plot cuts of an EPI image
mean_epi = mean_img('./output/synth_stripped_flirt_T1MNI.nii.gz', copy_header=True)
plot_epi(mean_epi, colorbar=True, cbar_tick_format="%i", title="mean_epi")
<nilearn.plotting.displays._slicers.OrthoSlicer at 0x7f35c3112900>
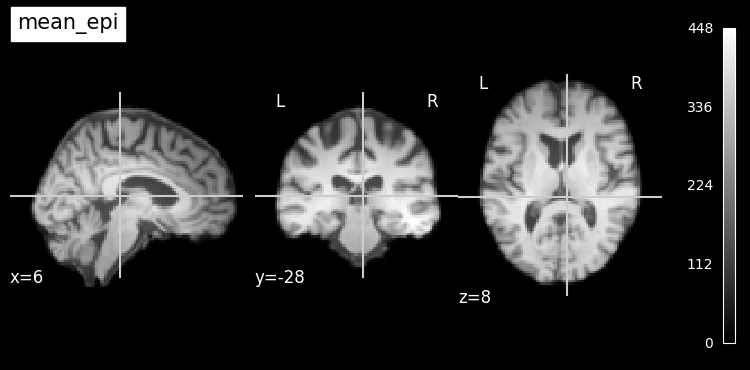
plot_anat and add_edges to see the overlay between two images#
# Check registration by overlaying anatomical image as edges (red) on top of mni template
t1_mni = './output/synth_stripped_flirt_T1MNI.nii.gz'
disp_edges = plot_anat(title="add_edges") #default=MNI152TEMPLATE
disp_edges.add_edges(t1_mni, color="r")
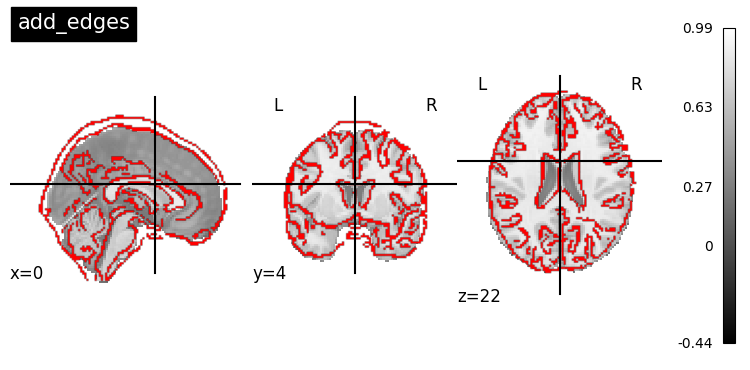
plot_roi#
Plot cuts of an ROI/mask image
plot_roi('./output/synth_mask.nii.gz', bg_img=t1_nipype,
title='Brain mask on skull-stripped brain')
<nilearn.plotting.displays._slicers.OrthoSlicer at 0x7f3565cbf610>
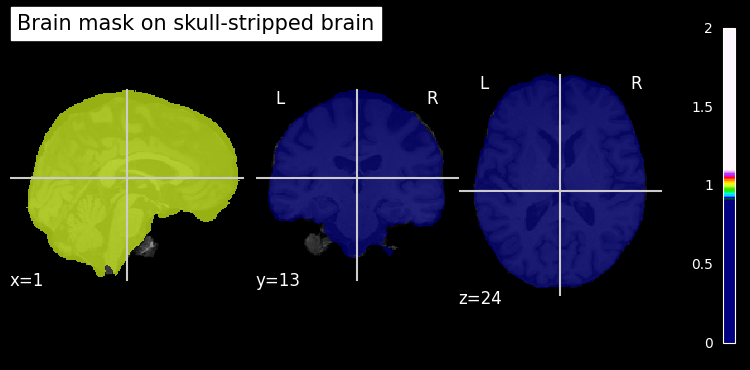
view_img#
Interactive html viewer
view_img(t1_nipype, bg_img='./ds001226/sub-CON02/ses-preop/anat/sub-CON02_ses-preop_T1w.nii.gz',
symmetric_cmap=False)
/opt/conda/lib/python3.13/site-packages/numpy/_core/fromnumeric.py:867: UserWarning: Warning: 'partition' will ignore the 'mask' of the MaskedArray.
a.partition(kth, axis=axis, kind=kind, order=order)
