Basic Nipype#
Author: Steffen Bollmann
Date: 17 Oct 2024
License:
Note: If this notebook uses neuroimaging tools from Neurocontainers, those tools retain their original licenses. Please see Neurodesk citation guidelines for details.
Citation and Resources:#
Dataset from OSF#
Shaw, T., & Bollmann, S. (2020). Dataset for Towards Optimising MRI Methods for ChAracterisation of Tissue (TOMCAT) [Data set]. OSF. https://doi.org/10.17605/OSF.IO/BT4EZ
Tools included in this workflow#
Nipype:
Esteban, O., Markiewicz, C. J., Burns, C., Goncalves, M., Jarecka, D., Ziegler, E., Berleant, S., Ellis, D. G., Pinsard, B., Madison, C., Waskom, M., Notter, M. P., Clark, D., Manhães-Savio, A., Clark, D., Jordan, K., Dayan, M., Halchenko, Y. O., Loney, F., … Ghosh, S. (2025). nipy/nipype: 1.8.6 (1.8.6). Zenodo. https://doi.org/10.5281/zenodo.15054147
FSL:
M. Jenkinson, C.F. Beckmann, T.E. Behrens, M.W. Woolrich, S.M. Smith. FSL. NeuroImage, 62:782-90, 2012
Smith S. M. (2002). Fast robust automated brain extraction. Human brain mapping, 17(3), 143–155. https://doi.org/10.1002/hbm.10062
AFNI:
Cox RW (1996). AFNI: software for analysis and visualization of functional magnetic resonance neuroimages. Comput Biomed Res 29(3):162-173. doi:10.1006/cbmr.1996.0014 https://pubmed.ncbi.nlm.nih.gov/8812068/
RW Cox, JS Hyde (1997). Software tools for analysis and visualization of FMRI Data. NMR in Biomedicine, 10: 171-178. https://pubmed.ncbi.nlm.nih.gov/9430344/
SPM12:
Karl J. Friston, K. J., Holmes, A. P., Worsley, K. J., Poline, J.-B., Frith, C. D., & Frackowiak, R. S. J. (1994). Statistical parametric maps in functional imaging: A general linear approach. Human Brain Mapping, 2(4), 189–210. https://doi.org/10.1002/hbm.460020402
Friston, K. J. (2007). Statistical parametric mapping: The analysis of functional brain images (1st ed). Elsevier / Academic Press.
Introduction#
This notebook provides a brief hands-on demonstration of Nipype, showing how it can be used to call neuroimaging tools directly from Python. For a more in-depth explanation of Nipype’s concepts and architecture, refer to the Nipype on Neurodesk.
Here we apply two common preprocessing steps to a 7T anatomical MRI image: skull stripping using FSL’s BET and spatial normalization to MNI space using SPM’s Normalize12, both called through Nipype’s interface system. We also use AFNI’s Edge3 to extract brain edges for visualization purposes.
Import Python modules#
%%capture
!pip install nibabel numpy scipy
from nipype.interfaces import fsl
from nipype.interfaces import afni
import nipype.interfaces.spm as spm
from ipyniivue import NiiVue
import matplotlib.pyplot as plt
import nibabel as nib
from matplotlib import transforms
from scipy import ndimage
import numpy as np
Demonstrating the module system in Python and Nipype#
# we can use module to load fsl in a specific version
import module
await module.load('fsl/6.0.4')
await module.load('afni/26.0.07')
await module.load('spm12/r7771')
await module.list()
['fsl/6.0.4', 'afni/26.0.07', 'spm12/r7771']
# test if fsl is loaded using any fsl command
!bet
Usage: bet <input> <output> [options]
Main bet2 options:
-o generate brain surface outline overlaid onto original image
-m generate binary brain mask
-s generate approximate skull image
-n don't generate segmented brain image output
-f <f> fractional intensity threshold (0->1); default=0.5; smaller values give larger brain outline estimates
-g <g> vertical gradient in fractional intensity threshold (-1->1); default=0; positive values give larger brain outline at bottom, smaller at top
-r <r> head radius (mm not voxels); initial surface sphere is set to half of this
-c <x y z> centre-of-gravity (voxels not mm) of initial mesh surface.
-t apply thresholding to segmented brain image and mask
-e generates brain surface as mesh in .vtk format
Variations on default bet2 functionality (mutually exclusive options):
(default) just run bet2
-R robust brain centre estimation (iterates BET several times)
-S eye & optic nerve cleanup (can be useful in SIENA - disables -o option)
-B bias field & neck cleanup (can be useful in SIENA)
-Z improve BET if FOV is very small in Z (by temporarily padding end slices)
-F apply to 4D FMRI data (uses -f 0.3 and dilates brain mask slightly)
-A run bet2 and then betsurf to get additional skull and scalp surfaces (includes registrations)
-A2 <T2> as with -A, when also feeding in non-brain-extracted T2 (includes registrations)
Miscellaneous options:
-v verbose (switch on diagnostic messages)
-h display this help, then exits
-d debug (don't delete temporary intermediate images)
Download test data#
%%bash
if [ -f ./sub-01_ses-01_7T_T1w_defaced.nii ]; then
echo "nii Output file exists, not downloading or unpacking again"
else
if [ ! -f ./sub-01_ses-01_7T_T1w_defaced.nii.gz ]; then
echo "nii.gz does not exist. So, it needs to be downloaded."
osfURL="osfstorage/TOMCAT_DIB/sub-01/ses-01_7T/anat/sub-01_ses-01_7T_T1w_defaced.nii.gz"
echo "downloading now ..."
osf -p bt4ez fetch $osfURL ./sub-01_ses-01_7T_T1w_defaced.nii.gz
fi
if [ -f ./sub-01_ses-01_7T_T1w_defaced.nii.gz ]; then
echo "nii.gz exists. So, it needs to be unpacked and deleted"
echo "unpacking now ..."
gunzip ./sub-01_ses-01_7T_T1w_defaced.nii.gz
fi
fi
nii Output file exists, not downloading or unpacking again
%ls
AA_Neurodesk_demo_tour.ipynb nipype_short.ipynb
MRIQC.ipynb papermill-slurm-submission-example.ipynb
Magic_commands.ipynb pydra_preproc_ants.ipynb
PyBIDS.ipynb pyscript.m
RISE_slideshow.ipynb pyscript_normalize12.m
bids_conversion.ipynb sub-01_ses-01_7T_T1w_defaced.nii
intro.md sub-01_ses-01_7T_T1w_defaced_brain.nii.gz
ipyniivue_ipywidgets.ipynb sub-01_ses-01_7T_T1w_defaced_edges.nii
nextflow_neurodesk.ipynb wsub-01_ses-01_7T_T1w_defaced.nii
nipype_full.ipynb y_sub-01_ses-01_7T_T1w_defaced.nii
Preprocessing with Nipype#
Skull Stripping and Extracting Edges#
# Run FSL BET for skull stripping
btr = fsl.BET()
btr.inputs.in_file = './sub-01_ses-01_7T_T1w_defaced.nii'
btr.inputs.frac = 0.4 # Fractional intensity threshold - lower values = larger brain mask
btr.inputs.out_file = './sub-01_ses-01_7T_T1w_defaced_brain.nii'
res = btr.run()
# Extract edges for visualization
edge3 = afni.Edge3()
edge3.inputs.in_file = './sub-01_ses-01_7T_T1w_defaced.nii'
edge3.inputs.out_file = './sub-01_ses-01_7T_T1w_defaced_edges.nii'
edge3.inputs.datum = 'byte'
res = edge3.run()
260409-04:15:16,283 nipype.interface WARNING:
FSLOUTPUTTYPE environment variable is not set. Setting FSLOUTPUTTYPE=NIFTI
260409-04:15:25,533 nipype.interface INFO:
stderr 2026-04-09T04:15:25.533060:++ 3dedge3: AFNI version=AFNI_26.0.07 (Jan 24 2026) [64-bit]
260409-04:15:25,534 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752:** AFNI converts NIFTI_datatype=4 (INT16) in file /home/jovyan/workspace/books/examples/workflows/sub-01_ses-01_7T_T1w_defaced.nii to FLOAT32
260409-04:15:25,535 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752: Warnings of this type will be muted for this session.
260409-04:15:25,536 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752: Set AFNI_NIFTI_TYPE_WARN to YES to see them all, NO to see none.
260409-04:15:25,537 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752:*+ WARNING: If you are performing spatial transformations on an oblique dset,
260409-04:15:25,538 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752: such as /home/jovyan/workspace/books/examples/workflows/sub-01_ses-01_7T_T1w_defaced.nii,
260409-04:15:25,539 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752: or viewing/combining it with volumes of differing obliquity,
260409-04:15:25,539 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752: you should consider running:
260409-04:15:25,540 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752: 3dWarp -deoblique
260409-04:15:25,541 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752: on this and other oblique datasets in the same session.
260409-04:15:25,542 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752: See 3dWarp -help for details.
260409-04:15:25,543 nipype.interface INFO:
stderr 2026-04-09T04:15:25.534752:++ Oblique dataset:/home/jovyan/workspace/books/examples/workflows/sub-01_ses-01_7T_T1w_defaced.nii is 1.253358 degrees from plumb.
260409-04:15:32,812 nipype.interface INFO:
stderr 2026-04-09T04:15:32.812676:** ERROR: output dataset name 'sub-01_ses-01_7T_T1w_defaced_edges.nii' conflicts with existing file
260409-04:15:32,814 nipype.interface INFO:
stderr 2026-04-09T04:15:32.812676:** ERROR: dataset NOT written to disk!
%ls
AA_Neurodesk_demo_tour.ipynb nipype_short.ipynb
MRIQC.ipynb papermill-slurm-submission-example.ipynb
Magic_commands.ipynb pydra_preproc_ants.ipynb
PyBIDS.ipynb pyscript.m
RISE_slideshow.ipynb pyscript_normalize12.m
bids_conversion.ipynb sub-01_ses-01_7T_T1w_defaced.nii
intro.md sub-01_ses-01_7T_T1w_defaced_brain.nii.gz
ipyniivue_ipywidgets.ipynb sub-01_ses-01_7T_T1w_defaced_edges.nii
nextflow_neurodesk.ipynb wsub-01_ses-01_7T_T1w_defaced.nii
nipype_full.ipynb y_sub-01_ses-01_7T_T1w_defaced.nii
# View 3D data
def view_slices_3d(image_3d, slice_nbr, vmin, vmax, title=''):
# print('Matrix size: {}'.format(image_3d.shape))
fig = plt.figure(figsize=(15, 4))
plt.suptitle(title, fontsize=10)
plt.subplot(131)
plt.imshow(np.take(image_3d, slice_nbr, 2), vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Axial');
plt.subplot(132)
image_rot = ndimage.rotate(np.take(image_3d, slice_nbr, 1),90)
plt.imshow(image_rot, vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Coronal');
plt.subplot(133)
image_rot = ndimage.rotate(np.take(image_3d, slice_nbr, 0),90)
plt.imshow(image_rot, vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Sagittal');
cbar=plt.colorbar()
def get_figure():
"""
Returns figure and axis objects to plot on.
"""
fig, ax = plt.subplots(1)
plt.tick_params(top=False, right=False, which='both')
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
return fig, ax
# load data
brain_full = nib.load('./sub-01_ses-01_7T_T1w_defaced.nii').get_fdata()
brain = nib.load('./sub-01_ses-01_7T_T1w_defaced_brain.nii.gz').get_fdata()
edges = nib.load('./sub-01_ses-01_7T_T1w_defaced_edges.nii').get_fdata()
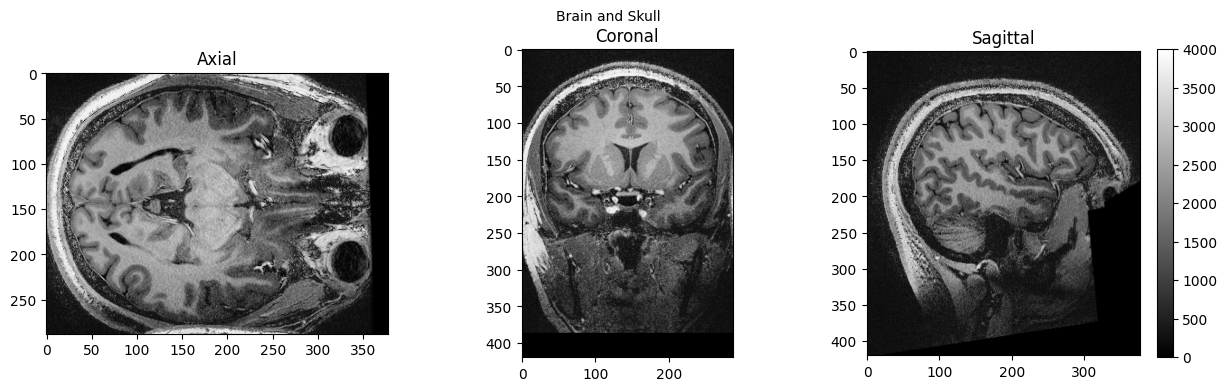
view_slices_3d(brain_full, slice_nbr=230, vmin=0, vmax=4000, title='Brain and Skull')
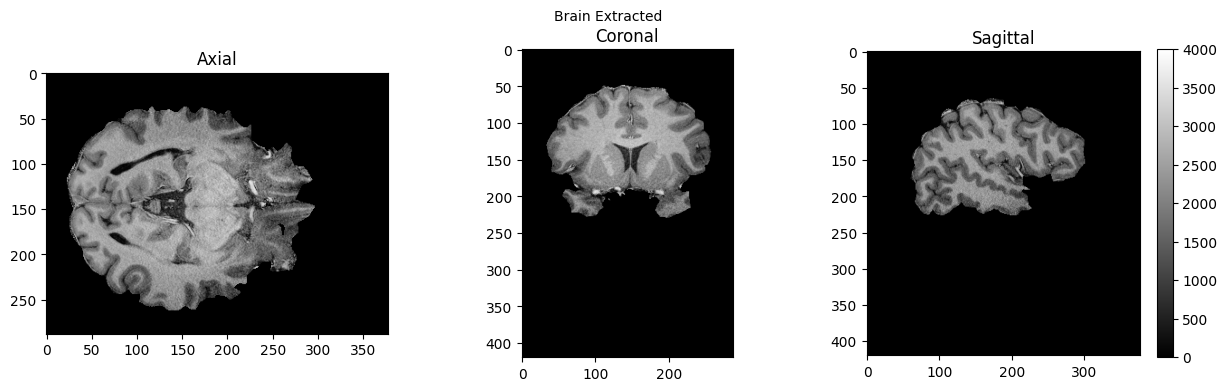
view_slices_3d(brain, slice_nbr=230, vmin=0, vmax=4000, title='Brain Extracted')
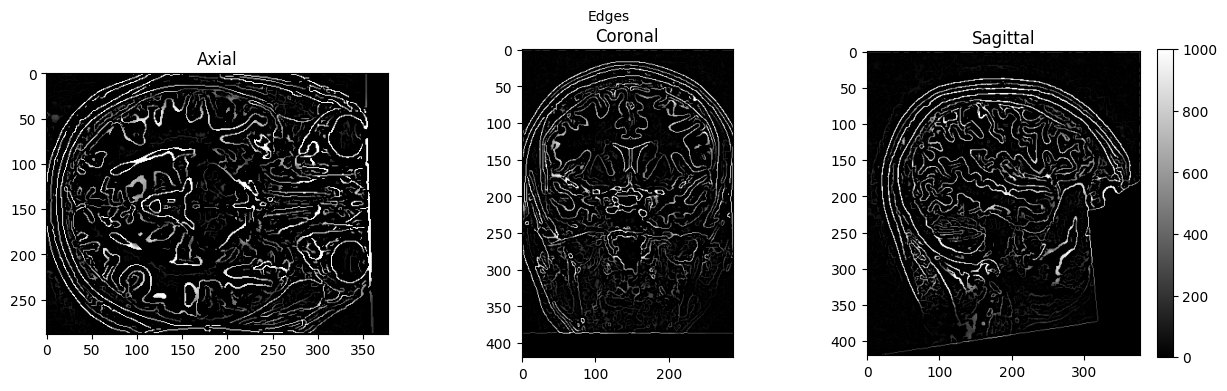
view_slices_3d(edges, slice_nbr=230, vmin=0, vmax=1000, title='Edges')
# Interactive exploration of the brain extracted and defaced brain
nv = NiiVue()
nv.load_volumes([{"url": "https://huggingface.co/datasets/neurodeskorg/neurodeskedu/resolve/main/data/examples/workflows/nipype_short/sub-01_ses-01_7T_T1w_defaced_brain_5268908826e9.nii.gz"}])
nv
MNI Normalization#
SPM can also be used in such a workflow, but unfortunately, this will trigger a warning “stty: ‘standard input’: Inappropriate ioctl for device”, which you can ignore (or help us to find out where it comes from):
# Normalize the anatomical image to MNI space using SPM Normalize12
norm12 = spm.Normalize12()
norm12.inputs.image_to_align = './sub-01_ses-01_7T_T1w_defaced.nii'
norm12.run()
stty: 'standard input': Inappropriate ioctl for device
stty: 'standard input': Inappropriate ioctl for device
<nipype.interfaces.base.support.InterfaceResult at 0x7fea37f2b100>
brain_full = nib.load('./wsub-01_ses-01_7T_T1w_defaced.nii').get_fdata()
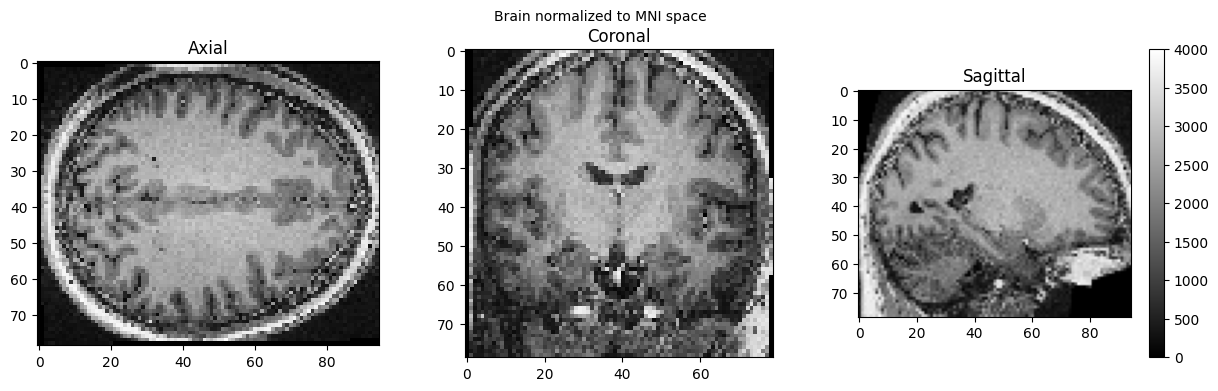
view_slices_3d(brain_full, slice_nbr=50, vmin=0, vmax=4000, title='Brain normalized to MNI space')
# Interactive inspection of the normalized brain with ipyniivue
nv = NiiVue()
nv.load_volumes([{"url": "https://huggingface.co/datasets/neurodeskorg/neurodeskedu/resolve/main/data/examples/workflows/nipype_short/wsub-01_ses-01_7T_T1w_defaced_b98f794d0a0a.nii"}])
nv
Dependencies in Jupyter/Python#
Using the package watermark to document system environment and software versions used in this notebook, alongside the Neurodesktop version extracted from the
JUPYTER_IMAGEorNEURODESKTOP_VERSIONenvironment variables.
import os
%load_ext watermark
%watermark
%watermark --iversions
neurodesktop_version = (
os.environ.get('JUPYTER_IMAGE', '').split(':')[-1] or
os.environ.get('NEURODESKTOP_VERSION', 'unknown')
)
print(f"Neurodesktop version: {neurodesktop_version}")
Last updated: 2026-04-09T04:18:24.441294+00:00
Python implementation: CPython
Python version : 3.13.9
IPython version : 9.7.0
Compiler : GCC 14.3.0
OS : Linux
Release : 5.15.0-171-generic
Machine : x86_64
Processor : x86_64
CPU cores : 32
Architecture: 64bit
ipyniivue : 2.4.4
matplotlib: 3.10.8
nibabel : 5.3.3
nipype : 1.10.0
numpy : 2.3.5
scipy : 1.16.3
Neurodesktop version: 2025-12-20
